Chitosan is derived from chitin, a linear polysaccharide that composes the hard shell of crustaceans (crabs, shrimp, etc.). After a degree of deacetylation of the chitin, chitosan can be dissolved in water and in acidic solutions. Chitosan has found many uses such as water purification, thickeners and stabilizers for food and pharmaceutical formulations, fertilizer, crop-immunizer and antihemorrhagic agents. Since chitosan is biodegradable and readily available, it should find even more uses as the demand for greener products grows.
The application requirements for various molecular weights of chitosan and various degrees of deacetylation makes accurate product characterization important. In this communication three different chitosans were measured to assess their molecular weight distribution and structure.
A Viscotek Size Exclusion Chromatography (SEC) system was used to characterize the molecular weight and the structure of chitosans samples without the need for column calibration. The front end of the system was a Viscotek GPCmax that included an online eluent degasser, a low pulsation isocratic pump and a variable injection volume autosampler. The system was controlled by the OmniSEC software that was also used to collect the data from the detectors. The detectors are all incorporated in the Viscotek TDA system along with the column compartment, keeping isothermal conditions for the injected material for ultimate reproducibility and stability. The TDA included light scattering detection at 90° and 7° angles, RALS and LALS respectively. The differential refractometer (RI) was in series with the LS detector, the last detector inline was the differential viscometer.
These detectors being in series allowed for the full sample to be characterized without splitting the flow. The OmniSEC software calculated the absolute molecular weight of the chitosan eluting using the low angle light scattering detector and the concentration using the RI detector. Finally the viscosity was measured for each data slice by the viscometer and the intrinsic viscosity calculated by combining with the concentration data. This triple detection system allowed the full characterization of the chitosan molecular weight distribution and viscosity distribution, allowing structural information to be determined.
The raw data from the triple chromatogram of the chitosan sample 2 is showed in figure 1. The red line is the RI signal which is proportional to the sample concentration. The black signal is the low angle light scattering signal (LALS) that is proportional to the molecular weight and the concentration. In SEC, the larger molecules exit the columns at a lower elution volume, so it is expected that larger chitosan will elute before lower molecular weights species, hence the skewing of the LALS chromatogram toward lower elution volume as the light scattering detector is more sensitive to the higher molecular weights. It should be noted that all 3 chromatograms are automatically adjusted for the interdetector delay and band broadening by the OmniSEC software. The blue chromatogram in figure 1 is from the differential pressure measurement in the bridge viscometer. The signal is proportional to the viscosity and the concentration. In this case the signal overlays well on the light scattering signal and is expected to show a well behaved polymeric distribution and structure. It should be noted that all 3 signals shows very little noise, ensuring a good measurement.

|
Three chitosan samples were measured. The absolute weight average molecular weight is given in table 1 for all 3 samples. Sample 1 and 3 have lower average molecular weight than sample 2. The OmniSEC software uses the concentration and the intensity of light scattered during the elution to calculate the molecular weight of each slice, reconstructing the weight average molecular weight distribution. The viscometer signal is used by OmniSEC similarly to obtain the intrinsic viscosity of the 3 chitosans. Here also, sample 1 and 3 have lower viscosity than sample 2. Finally the Stokes-Einstein equation is used to calculate the average hydrodynamic radius of the samples using the 3 detectors. It should be noted that the average is weighted by the polymer weight, not by the intensity ("Z-average") as in uncorrected dynamic light scattering measurements. As expected the sample 1 and 3 are smaller on average than sample 2.
| Sample | Mw (g/mol) | [η] (dL/g) | Rh (nm) |
|---|---|---|---|
| Chi1 | 44,411 | 1.32 | 9.2 |
| Chi2 | 104,326 | 2.72 | 15.5 |
| Chi3 | 48,989 | 1.50 | 10.1 |
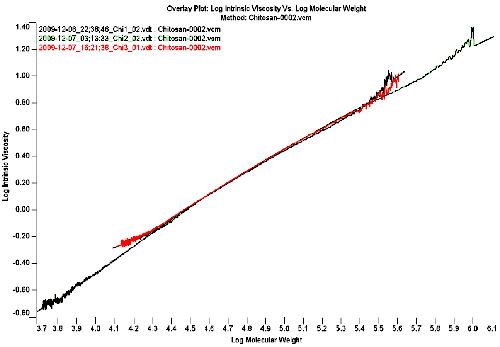
Figure 2 shows the overlaid Mark-Houwink plot if the 3 chitosan samples. The Mark-Houwink plot is constructed by taking the intrinsic viscosity and plotting it against the molecular weight for each slice on a log-log scale. Sample 1 in black has the lowest molecular weight fraction as it is extending the most to the left, sample 2 in green has the largest chitosan chains as it is extending the most to the right of the graph. Sample 3 in red has the lowest polydispersity as the polymer chain doesn't exhibit such a large span of molecular weights.
|
The most important information obtained from the Mark-Houwink plot in figure 2 is the fact that all 3 samples superimpose, indicating that the structure of the polymer backbones are similar for all 3 chitosans, despite the different molecular weights. The linear graph also indicates that the polymers are linear polymers and there is no branching in these chitosan samples.
The Viscotek triple detection system has been used successfully to characterize the 3 chitosan samples, showing that the polymer structures are similar despite the different molecular weight distributions. The molecular size of the polymer has been measured precisely using triple detection. The linear Mark-Houwink plot indicates a linear polymer without any branching. This technique is equally suitable for other polysaccharides and, indeed, for all synthetic or natural polymers.