Use DSC to select antibody with lowest propensity to aggregate. An important consideration during the engineering process of an antibody is both the inherent conformational stability of the antibody as well as long-term stability. Understanding these stability parameters and what trends they follow will be critical throughout the biotherapeutic development. Differential scanning calorimetry (DSC) is a potent tool in probing these qualities. As a stability indicating assay, DSC provides insights for the selection of stable protein construct candidates. In the case of multi-domain proteins, DSC uniquely allows the investigation of the stability of individual domains from multi-domain proteins like antibodies and Fc-conjugated proteins. In this application note, we compare stability data predicted by DSC of different engineered antibodies with SEC-HPLC data from accelerated stability studies. For the engineered proteins studied, there is a correlation between decreased thermostabilty based on DSC data, and greater aggregation formation during accelerated stability studies. This study demonstrates how DSC can be a useful tool in stability screening of engineered proteins.
Whenever a biotherapeutic protein, like a monoclonal antibody, is engineered and expressed in a recombinant system, it is important that the engineered protein maintains its desired potency, efficacy, stability, and solubility. During the engineering process, various protein characteristics are analyzed and optimized, such as:
At Applied Molecular Evolution, protein engineering is performed by directed evolution technology (Figure 1) which permits simultaneous optimization of multiple protein properties. The screening method, rather than the traditional selection method, can solve more complex problems, and captures more structure/function data.
|
One of analytical/physicochemical methods employed during the course of protein engineering at AME is Differential Scanning Calorimetry (DSC) to compare the thermostability of parent and engineered antibodies. DSC is commonly employed to assess the thermal and conformational stability of a protein. The melting temperature of a protein, or of individual domains, can be obtained from the DSC profile, and if the reaction is reversible the thermodynamic parameters of the unfolding can be determined. A protein is considered more thermostable when the Tm (transition temperature) is higher. When thermostability data are compared to other bioanalytical methods, like size exclusion chromatography (SEC), others have observed a correlation between greater thermostability (higher Tm) and decreased aggregation (From data presented by at the 2007 Trends in Calorimetry Conference). Antibodies with a higher Tm and decreased aggregation formation have the potential for improved long-term stability thus potentially making better biotherapeutics.
In this manuscript we compare the stability of three different engineered antibodies as determined by SEC-HPLC analysis during accelerated stability studies, with the stability predicted by DSC. The results demonstrate a correlation between decreased thermostabilty based on DSC data, and greater aggregation formation during accelerated stability studies for the antibodies studied. Thus, the results demonstrate how DSC can be a useful tool in stability screening of engineered proteins.
Antibody samples were dialyzed against 10 mM citrate buffer at pH 6.5 and diluted to 1 mg/mL. Scans were performed using an automated MicroCal VP-Capillary DSC equipped with a 96-well plate autosampler. Scans ran from 20-95°C at 200°C/h and were analyzed using Origin 7.0. The unfolding transitions of each antibody were fit using the non-two-state unfolding model.
For accelerated stability study, the same set of antibody samples described above was stored at 60°C for up to 5 days. Aliquots were withdrawn every day and centrifuged at 15,000 g for 20 minutes to remove precipitated antibodies. The supernatant was used for determining soluble protein concentration and the percentage of soluble high molecular weight aggregates. Analytical size-exclusion chromatography (SEC) was performed using a Tosoh G3000SWXL (300 X 4.6 mm) column at a flow rate of 0.5 mL/min.
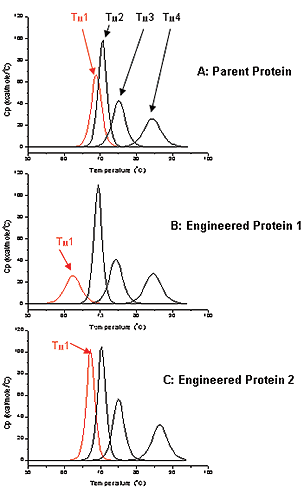
While the functional and biological properties of antibodies are well known, little has been published concerning the folding and stability of the many Ig-Fold domains of antibodies within the context of the full-length molecule. DSC can be used to investigate multi-domain folding thermodynamics to guide antibody design. In this case study, the parent antibody that was used as the starting point for engineering has four distinct unfolding domains in the DSC thermogram (Figure 2A). Each unfolding domain has a measurable transition temperature (Tm1 to Tm4). Tm1 was monitored for thermostability for this protein. For the parent protein Tm1 was approximately 69 °C (Figure 2A).
In this study, antibodies were being engineered for improved potency, and after the first round of screening, we chose Antibody 1 for further characterization in terms of biophysical properties. DSC data showed that Tm1 of Antibody 1 was approximately 62 °C (Figure 2B), or 7 degrees lower when compared to the Tm1 of the parent antibody, suggested that Antibody 1 had a decreased thermostability relative to the parent protein.
A second round of engineering and screening, resulted in Antibody 2. This particular antibody has Tm1 of about 69 °C, shifting up about 7 degrees relative to Tm1 for Engineered Antibody 1, and having a Tm comparable to the parent protein (Figure 2C). This result suggests that Antibody 2 have comparable thermostability, compared to parent antibody. Since proteins with greater thermostability may be more stable under accelerated stability conditions, a potential advantage is a longer shelf life of the biotherapeutic.
|
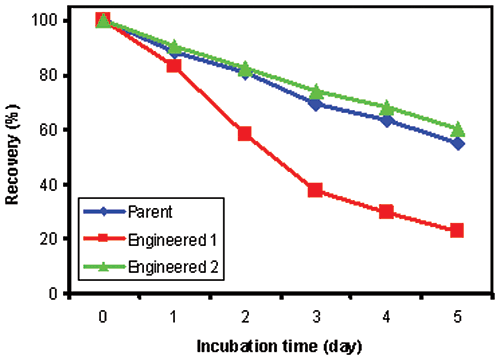
Since the parent and engineered antibodies showed comparable stability at 37°C (data not shown), we chose to perform accelerated stability studies at 60°C, which was close to Tm1 of Antibody 1. Figure 3 shows the percent recovery of soluble protein after storage at 60°C for indicated number of days. Parent and Antibody 2 were both very stable at 60 °C, and had comparable protein recovery. However, Antibody 1 has a significant decrease in recovery after five days of storage at 60 °C. The stability predictions made by DSC of the three antibodies tested correlate well with measurements of percent recovery of soluble protein from accelerated stability studies. In comparison, since DSC forces the unfolding event(s), stability information can be assessed more quickly.
|
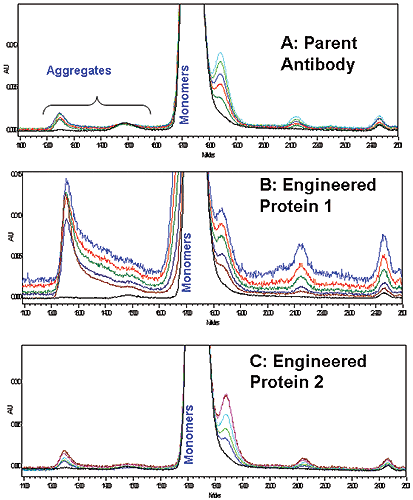
Accelerated stability studies at 60°C also indicated that parent and Antibody 2 had comparable, low amount of soluble aggregate formation, based on SEC-HPLC data (Figure 4A and 4C). The same experiment performed with Antibody 1 showed greater soluble aggregate formation, compared to parent and engineered antibody 2 (Figure 4B). Stability predictions made with DSC correspond well to what is predicted by longer-term SEC-HPLC aggregation studies for the antibodies tested, thus microcalorimetry may be a useful tool for rapidly screening protein stability in solution.
|
Even though Antibody 1 has a higher potency than the parent antibody, accelerated stability studies suggest that this protein has increased soluble and insoluble aggregation, compared to the parent antibody and a second engineered antibody. Greater aggregation formation on accelerated studies suggests potential long-term stability and shelf life issues. Since Antibody 2 has a higher Tm and improved stability in accelerated studies, this protein may be the better choice as a biotherapeutic. Further studies are needed to correlate improved stability in accelerated stability studies and improved long-term stability and longer shelf life of this engineered antibody.
DSC was used to study the thermal stability of antibodies that were being engineered for improved potency. In this study, we demonstrated a correlation between decreased thermal stability based on DSC data, and greater aggregation formation during accelerated stability studies (at 60°C). The SEC-HPLC stability data agree quite well with the stability predicted by DSC, suggesting that the thermal stability data obtained from DSC correlates with protein stability. Information derived from DSC can indicate potential long-term stability issues, thus making the technique a useful tool in the screening and selection of engineered proteins.