Combining dynamic light scattering (DLS) and Raman spectroscopy provides the ability to extract a wealth of chemical, structural, and physical parameters from proteins at both low and formulation concentrations. DLS, used to study protein size, and Raman, used to determine protein conformation, is combined into a single instrument to monitor stressed protein samples under identical conditions.
DLS is ideally suited to measuring the size of proteins at low concentrations (down to 0.1 mg/mL), while Raman spectroscopy is ideally suited to deriving secondary and tertiary structural markers from higher concentration protein samples (50 mg/mL or greater). One practical area of interest is understanding the lowest concentration of a protein that can be studied using these combined analytical techniques. Because DLS is known to be appropriate for analyzing low concentration samples, this note will outline the lowest practical protein concentration limit for acquiring Raman spectra, and determine if the spectra provide useful secondary and tertiary structure information. Results using lysozyme indicate that useful information from concentrations as low as 3 mg/mL can be acquired, while monoclonal antibody samples (mAbs) may require a higher concentration to obtain correct structural and thermal information.
The ability of a combined DLS/Raman instrument to study various concentrations of protein samples is explored. DLS, with its capability to measure protein size, is ideal for low concentration samples, while Raman, with its ability to interrogate protein secondary and tertiary structure, is ideal for high concentration samples. Joining the techniques into a single technology allows for the determination of size and structure on a single small volume sample. However, because the complementary techniques have differing ideal concentrations, it is important to first establish the upper and lower practical concentration boundaries for a protein sample. This note focuses on determining the lowest concentration limit for studying proteins with combined DLS/Raman.
Malvern Instruments’ Zetasizer Helix (ZS Helix) integrates a fiber-coupled Raman spectrometer with a Zetasizer Nano ZSP to provide DLS (colloidal stability) and Raman (conformational stability) data sequentially on a single sample. The Zetasizer Nano system integrates proprietary non-invasive backscatter (NIBS) detector technology with dynamic (DLS), static (SLS) and electrophoretic (ELS) light scattering to measure the hydrodynamic radius of proteins from 0.15 nm - 5 µm, from 0.1 mg/mL to ≥ 100 mg/mL. Raman spectra are collected using 785 nm excitation (~280 mW) from 150 cm-1 - 1925 cm-1 at 4 cm-1 resolution. Sample aliquots (~120 µL) are placed into a 3 mm quartz cuvette and positioned in a temperature controlled compartment that provides temperature control from 0°C - 90°C ± 0.1°C. Thermal ramp studies are conducted by collecting Raman and DLS data over a series of pre-defined 0.1°C - 5°C step-wise increments. Isothermal incubation studies are conducted by collecting a series of Raman and DLS data over a pre-defined time interval at a desired temperature set-point.
Raman spectral data collection conditions are optimized for low concentration protein samples to determine a practical detection limit for acquiring Raman spectra for protein therapeutics. Two samples are analyzed, lysozyme and a mAb, at multiple concentrations. Secondary structural markers, including amide I (1600 cm-1- 1700 cm-1) and amide III (1200 cm-1 - 1350 cm-1), are observed, along with tertiary structural markers, such as tyrosine (Tyr) at ~830 cm-1 and tryptophan (Trp) at ~1550 cm-1 during thermal ramp experiments. Changes in these markers indicate a change in the structure of the protein during heating, so the ability to detect any changes is inherently important to studying protein structure.
This application note focuses on determining the lowest concentration of protein that can be studied using a combined DLS/Raman instrument. Because DLS is ideal for lower concentration samples (addressed in a previous application note), most of the effort here has been employed in optimizing the Raman acquisition parameters. Up to a certain limit (saturating the detector), there is a positive linear relationship between integration time and Raman signal, and a square root relationship between the number of co-adds and the Raman signal-to-noise. Therefore, to increase the Raman signal from lower concentration samples that have less Raman scattering particles available, it makes sense to increase the integration time as long as possible and increase the number of co-additions of multiple scans (co-adds). However, in an effort to keep experiment times practical, all acquisition times are kept at or below 30 minutes for both the lysozyme and mAb low concentration samples. The higher concentration samples are used to establish expected changes in the protein structure. Below is a summary of the acquisition parameters used for collecting both lysozyme and mAb data (Table 1).
| Sample | Integration time (s) | Co-adds (#) | Raman acquisition time (s)/T | Total acquisition time (min)/T |
|---|---|---|---|---|
| 35 mg/mL lysozyme | 15 | 10 | 150 | 5.5 |
| 12 mg/mL lysozyme | 30 | 10 | 300 | 8 |
| 3 mg/mL lysozyme | 40 | 40 | 1600 | 30 |
| 50 mg/mL mAb | 20 | 10 | 200 | 6 |
| 10 mg/mL mAb | 20 | 10 | 200 | 6 |
| 5 mg/mL mAb | 25 | 40 | 1000 | 20 |
Thermal ramping experiments are performed to interrogate 3 mg/mL, 12 mg/mL, and 35 mg/mL lysozyme samples in citrate buffer at pH 4. The highest concentration sample is used as a point of reference for the low concentration sample comparisons.
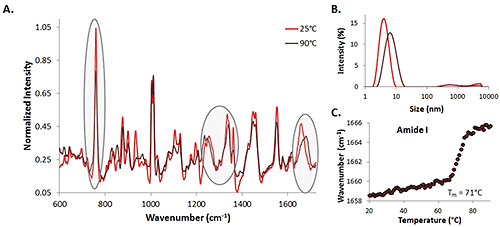
Spectra of a 35 mg/mL lysozyme solution at 25°C (represented by the light red trace) and 90°C (represented by the dark red trace) are included (Figure 1A). Some protein markers of interest are highlighted, indicating the changes in secondary and tertiary structure during the temperature ramp experiment. Changes are evident in the amide I (~1600 cm-1- 1700 cm-1), amide III (~1200 cm-1- 1350 cm-1), and the α-helix skeletal (930 cm-1 - 950 cm-1) regions, as well as the Trp at 760 cm-1.
DLS size data (intensity distribution) is also included for the high concentration lysozyme sample (Figure 1B). As expected, results indicate that as the temperature increases, the size of the lysozyme increases as well. This suggests protein unfolding (and possibly aggregation) is occurring as the temperature rises, which is consistent with the spectral data that indicates that the structure of the protein is changing as the temperature increases. Additionally, the center of mass (COM) shift with temperature for the amide I peak is included (Fig. 1C). As the temperature increases, the COM shifts to higher wavenumbers, indicating that the structure is changing from α-helix to β-sheet rich as the temperature increases. At this relatively high concentration, both the spectral and size transitions are obvious, the Raman data can be used for both protein detection and characterization, and transition information can be extracted. This information is used as a comparison for the lower concentration data.
|
To increase the signal to noise ratio of the Raman data for the lower concentration samples, longer integration times and more accumulations must be used than those needed for the higher concentration samples. This results in longer data collection time per spectrum. Data accumulation times as long as 30 min for a single temperature point (DLS and Raman) were used for the lowest concentration lysozyme samples (Table 1). Though this results in a longer experiment, it allows for the identification of secondary and tertiary structural markers in the Raman data, and in some cases, allows for the determination of spectral changes resulting from thermal stress.
Raman spectra, DLS intensity distributions, and amide I COM shifts for high (35 mg/mL), medium (12 mg/mL) and low (3 mg/mL) lysozyme concentrations are compared in Figure 2.
|
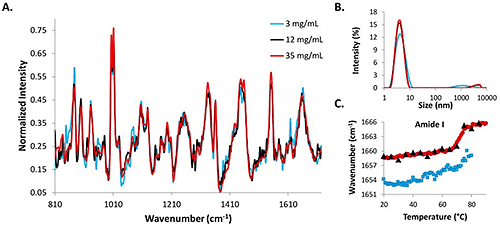
The Raman spectrum for the 35 mg/mL (red line) data is the least noisy, while the data from the 3 mg/mL concentration lysozyme sample (blue line) is the most noisy, and the 12 mg/mL sample (black line) falls somewhere in the middle of the two (Fig. 2A). Even with the increased acquisition times, the signal from the low concentration sample is noisier than those from the two higher concentration samples. Intensity distributions from the DLS data indicate that the protein molecules in solution are all ~4 nm at 25°C (Fig. 2B). The similarity between the size data and the spectral data at the starting temperatures for all three concentrations indicates that DLS/Raman can be used for lysozyme detection at concentrations as low as 3 mg/mL.
To further evaluate the ability of the DLS/Raman instrument to detect low concentration protein samples, the COM shift for the amide I peak is included for all concentrations (Fig. 2C). The red line represents the high concentration sample and clearly exhibits a shift to higher wavenumbers with a melting temperature of 71°C (determined by fitting a sigmoid to the curve). The 12 mg/mL data shows a similar transition (black triangles that are overlaying the red circles); however, melting temperature cannot be confirmed with this sample because temperature increments were 5°C steps instead of 1°C steps (to avoid possible kinetic effects). The similarity between the trends is apparent, though. The 3 mg/mL COM shift is less obvious and cannot be confirmed. Due to time required for the measurement, data was only collected until 80°C, as opposed to 90°C as with the higher concentration samples, and there is no sharp transition as was observed with the higher concentration samples. A more gradual transition could potentially be taking place at lower temperatures, but it is impossible to tell if this transition is the result of protein structural changes solely as the result of increased temperature, or if it is the result of the temperature increase in combination with the greater length of time during which the protein solution is exposed to higher temperatures (as a result of the long data acquisition times at each temperature increment).
Additional comparisons between the high concentration and low concentration samples are made by using an in-house PLS model to determine the secondary structural composition of 5 concentrations of lysozyme (Fig. 3).
|
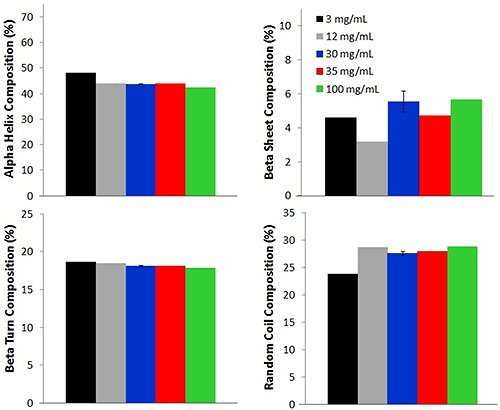
Four different secondary structure compositions: alpha helix, beta sheet, beta turn, and random coil, are predicted for each of the lysozyme samples. For all concentrations, the alpha helix percentage is the highest, with an average percentage of 44.0% ± 2.0%. This value agrees with literature reports of lysozyme solutions at pH values near 4 [1,2]. Random coil composition, 28.0% ± 2.0%, is reported as the second highest concentration, followed by beta turn (18.0% ± 0.3%), and beta sheet (6.0% ± 1.0%). Although a small difference (reported as standard deviation) is observed between the various concentrations, the values are very similar across the concentrations, again indicating that the Raman data collected from the low concentration samples is a good representation of the lysozyme.
The results from the lysozyme thermal ramping studies indicate that concentrations as low as 3 mg/mL can be detected/identified using Raman spectroscopy, but for any information regarding the conformational changes during heating, a higher concentration sample is probably needed.
Similar to lysozyme, high concentration (50 mg/mL), medium concentration (10 mg/mL), and low concentration (5 mg/mL) samples of a mAb were studied using the combination of DLS/Raman. Thermal ramping experiments were performed from 25°C - 90°C with 0.2°C - 0.5°C steps for all concentrations. High concentration data was collected first and used as a comparison for the low concentration sample data.
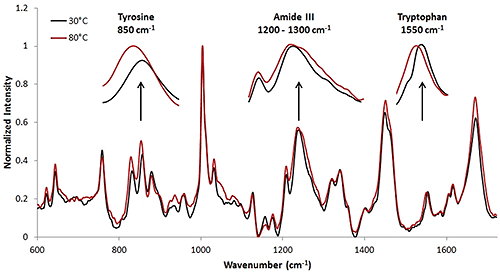
The Raman collection time for the high concentration sample was ~ 3.3 minutes per temperature increment (Table 1). This collection time resulted in a Raman spectrum with a high signal to noise ratio. The Raman spectra at 30°C and 80°C are included (Fig. 3). Here, the wavenumber shifts that indicate protein structural changes are less obvious than in the model protein, lysozyme, that was studied above. However, upon analysis, it is evident that tertiary structural changes are occurring when the protein solution is under thermal stress. In this case, the tertiary structural changes are more prominent than the secondary structural changes. Some of these secondary and tertiary structural shifts are highlighted.
|
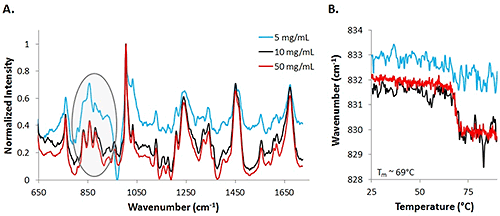
Assessing the mAb data at lower concentrations proved to be more difficult than analyzing the low concentration lysozyme sample, and more difficult even than the high concentration mAb sample. Figure 4 includes spectra from 50 mg/mL (red line), 10 mg/mL (black line), and 5 mg/mL (blue) mAb samples at 25°C. The highest concentration sample results in the least noisy spectrum, while the data from the low concentration sample is the most noisy. With the mAb samples, secondary structure does not change significantly, so small changes in the tertiary structure are evaluated. With the low concentration sample, it is much more difficult to identify the markers. One important area of interest is highlighted, and demonstrates the difficulty in seeing the markers, in this case, Tyr, in the 830 cm-1 - 855 cm-1 range. Results indicate that the low concentration data is noisier than the high concentration data, similar to the results obtained from the lysozyme samples.
|
To further establish that the data between the high and low concentration samples differs in quality, COM shifts are graphed for Tyr markers at 50 mg/mL, 10 mg/mL, and 5 mg/mL samples (Fig. 4B). A significant shift is observed for the 50 mg/mL solution (red line) and an estimated melting temperature of 69°C is obtained by fitting a sigmoid to the curve. The 10 mg/mL data also shows a similar shift with the increase in temperature (black line). However, by observing the COM data from the 5 mg/mL sample, there is not an obvious shift in the COM with an increase in temperature. The signal is dominated by the noise, even with an increase in integration time. Based upon the information obtained from the mAb sample at various concentrations, concentrations as low as 5 mg/mL can be used for detection of the sample, but for the elucidation of any type of thermal transitions, concentrations need to be at least as high as 10 mg/mL.
The combination of DLS/Raman can easily be used for the elucidation of protein conformation (Raman) and size (DLS) changes when exposed to stressors. The lower concentration limit for Raman detection, traditionally only used to study high concentration samples, has been evaluated here by performing thermal ramping experiments. A model protein, lysozyme, at concentrations as low as
3 mg/mL can be studied, with the identification of secondary and tertiary structural markers. Monoclonal antibodies, on the other hand, with their more stable secondary structures, are more likely to exhibit only tertiary structural changes. Some of these markers can be identified at concentrations as low as
5 mg/mL, but for any transition information, concentrations need to be as high as 10 mg/mL. The combination of these complementary techniques can allow for the understanding of protein unfolding and aggregation pathways at both low and high concentrations.
Malvern Instruments' Bioscience Development Initiative (BDI) was established to accelerate innovation, development and the promotion of new technologies, products and capabilities to address unmet measurement needs in the biosciences markets.