Dr.Stefano Bruno, Dr. Serena Faggiano, Dr Luca Ronda, University of Parma, Italy
The oligomeric state of globins is a crucial piece of information required to understand their structure-function relationships. It is known to vary significantly within the superfamily of proteins consisting of 1, 2, 4, 12, 24 globin subunits that have been found in nature. Within the group of globins, it is particularly critical to assess the size of hemoglobin-based blood substitutes, as this has been shown to correlate with the extent of both extravasation and renal filtration, two factors responsible for the toxicity of these products once they are administered to patients.
The assessment of the oligomeric state of globins is usually carried out by gel filtration chromatography, a fairly time consuming technique. Moreover, chromatography measurements imply dilution of the sample which can complicate the data interpretation as concentration-dependent dissociation of the subunits can occur.
Dynamic light scattering (DLS) is a rapid batch technique allowing the measurement of samples without the need for dilution, hence avoiding the problem of possible complex dissociation. The technique is non-invasive and the samples can be recovered after measurements for further use.
DLS measures the time dependent fluctuations in scattered light from molecules in a solution due to Brownian motion to determine the translational diffusion coefficient and subsequently determine the hydrodynamic radius (RH) from the Stokes-Einstein equation. The RH determined corresponds to the radius of a sphere diffusing with the same speed as the molecule in solution. The RH depends thus on the shape of the molecule, with an increased RH for elongated molecules compared to spherical molecules of the same molecular weight. For similarly shaped molecules, a calibration curve can be constructed to estimate the molecular weight based on the measured RH.
All globins share very similar 3D structures. In this application note, a calibration curve for globins has been constructed by measuring the following globins, where the molecular weight and subunit conformation is known: Myoglobin, Human hemoglobin, AHb1, AHb2 (two globins from Arabidopsis thaliana), rice Hb1 and Hemoglobin Polytaur (an auto-polymerizing recombinant globin proposed as a blood substitute). To extend the range of sizes measured, two other proteins were added: Ferritin from spinach and Hemocyanin from Eurypelma californicum, as they are high molecular weight proteins with a globular shape. Table 1 summarizes the molecular weight and number of subunits in each globin.
| Protein | No. of Subunits | Known Mw [kDa] | Peak Mean [nm] | %Peak Intensity | %Peak Volume |
|---|---|---|---|---|---|
| Myoglobin | 1 | 17.6 | 2.403 | 97.7 | 100 |
| AHb2 | 1 | 17.8 | 2.341 | 53.2 | 99.9 |
| Rice Hb1 | 2 | 37 | 2.985 | 90.9 | 100 |
| AHb1 | 2 | 36 | 3.016 | 80.8 | 100 |
| HbA | 4 | 64.5 | 3.305 | 100 | 100 |
| Polytaur | 12 | 186 | 6.778 | 88.3 | 99.8 |
| Ferritin | - | 440 | 8.74 | 82.8 | 99.9 |
| Hemocyanin | - | 1720 | 12.36 | 100 | 100 |
The resulting calibration curve was thereafter used to estimate the apparent molecular weight of two hemoglobin-based oxygen carriers (HBOCs). Both of them are PEGylated hemoglobins, one called PEGHboxy and mimicking the MP4 product (Hemospan®) from Sangart, and the other is called PEGHbdeoxy [2]. Both are hemoglobin molecules (64 kDa) conjugated with a number of 6-8 polyethylene glycol (PEG) chains (5.5 kDa).
All measurements were performed on the Zetasizer Nano ZS, a dynamic light scattering (DLS) instrument, using non-invasive backscatter (NIBS) detection.
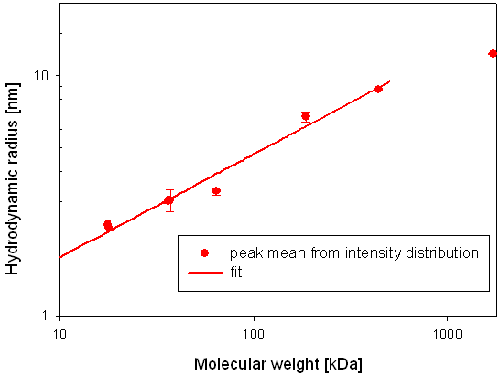
The calibration curve was constructed by plotting the measured hydrodynamic radius versus the known molecular weights. The data points were fitted with an equation of the following form RH = (a* Mw)^b [1] to obtain a relationship between the hydrodynamic radius and the molecular weight (Mw) for globins.
All protein concentrations were in the range 5-500 μM in saline buffer, where, for each protein, no subunit dissociation is predicted to take place.
All samples were filtered before measurement using 0.22mm or smaller pore size to remove large aggregates.
Most of the samples showed a multimodal correlation function indicating the presence of several populations in the samples.
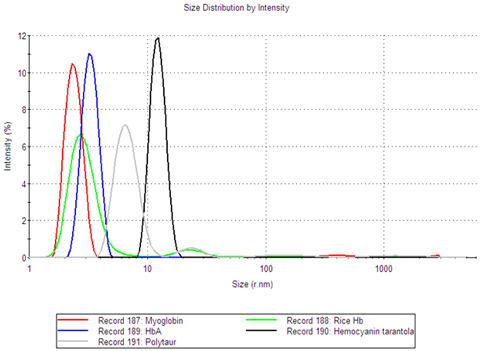
Although the samples are polydisperse, the size distribution analysis provides information about the individual populations and allows for the determination of the oligomeric state of the globin samples. Figure 1 shows the intensity size distributions for a number of the globin samples. The figure clearly shows that the different samples have multiple size peaks.
|
The fraction of the sample in each size peak was obtained from the volume size distributions, determined using Mie theory. It was shown that, for all globin samples, the smallest size peak was the main component of the sample, making up at least 99.8% of the sample volume (Table 1).
The size of the main component in each sample was therefore determined as the mean value of the smallest size peak. In Table 1, a summary of the peak statistics for the main peak of each sample is shown. It is noteworthy that proteins of similar molecular weight, such as Myoglobin (17.6kDa) and AHb2 (17.8kDa) have very similar hydrodynamic radii, 2.40nm and 2.34nm respectively. The same can be seen for Rice Hb1 (37kDa) and AHb1 (36kDa), where the RH was measured to be 2.99nm and 3.02nm, respectively.
These data confirm that it is possible to use DLS as a quick and user friendly screening tool to determine the oligomeric state of a globin sample, as well as a tool to detect the presence of aggregates in samples.
The hydrodynamic radius of the main component in each globin sample is plotted versus the known molecular weight of the samples in figure 2. The data fit, giving the relationship between molecular weight (Mw) and hydrodynamic radius (RH) has been added to figure 2.
|
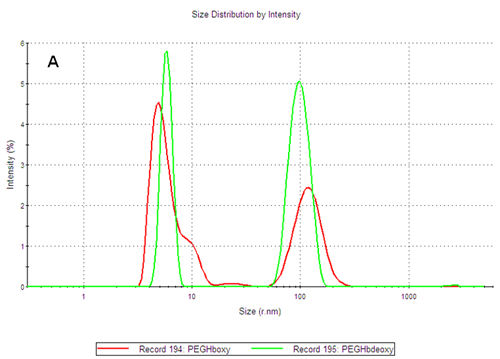
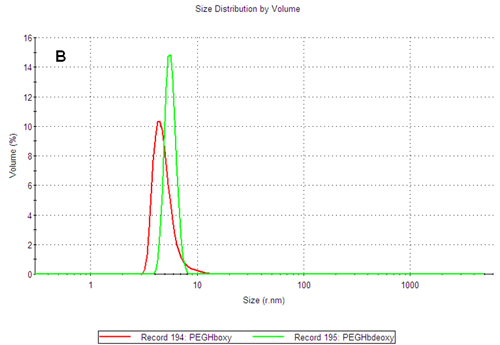
The PEGylated hemoglobins contain multiple populations, with the smaller size peak being the main component of both samples. Figure 3 shows the intensity and volume size distributions for the two samples. The two samples differ as the PEGHbdeoxy sample shows a narrow main size peak, with RH of 5.85nm, while the PEGHboxy sample contains at least two populations in its main size peak as indicated by a shoulder, with RH values corresponding to 4.89nm and ca 9nm respectively.
|
|
Using the calibration curve previously established, the estimated molecular weight of the main components of PEGHboxy are 108 kDa and 450kDa while the single peak of the PEGHbdeoxy sample corresponds to 165kDa. As the PEG chains conjugated to the protein affect the diffusion of the molecules, the calibration curve should only be used to estimate the corresponding weight of a globin with the same structure as the PEGylated hemoglobin products. The importance of measuring the size and thermal stability of these two products is discussed in more detail in another application note [2].
The size distributions of various globins have been determined using DLS. DLS allows the determination of the hydrodynamic radius of globin samples, and the identification of monomeric, dimeric and higher oligomers of globin samples. It has been shown that DLS can be effectively used to estimate the Mw and determine the oligomeric state and the fraction of aggregates present in globin samples.
By establishing a calibration curve for similarly shaped proteins, DLS was used to estimate the molecular weight of globins from their hydrodynamic radius, thus constituting a fast alternative or complement to gel filtration chromatography.
[1] Claes P., Dunford M., Kenney A., and Vardy P. " An on-line dynamic light scattering instrument for macromolecular characterization" in "Laser Light scattering in biochemistry" SE Harding, DB Sattelle, and VA Bloomfield (eds) pp 66-76, RSC (1992).
[2] "Size, oligomeric state and thermal stability of PEGylated hemoglobins", MRK1294-01
These results were obtained by Drs. Stefano Bruno, Serena Faggiano, and Luca Ronda at University of Parma, Italy with the help of Alice Brun, Alfatest srl, Italy.