The amount of UV light absorbed by a compound is an intrinsic property of compound's chemical composition. As analytical scientists, we exploit this property to monitor elution profiles from chromatography columns. The premise for using UV as a concentration detector is that UV absorbance increases with the amount of absorber in solution. In this document, we explain how absorbance, concentration and the corresponding coefficients are related and how dA/dc is used in the OmniSEC software.
At a given wavelength, the absorption of UV light by a compound is defined as the ratio of incident light (I0) to transmitted light (It). The relationship between this absorbance and the compound concentration and path length is described by the Beer-Lambert law (Equation 1).
Equation 1
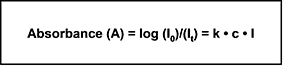
|
If absorbance is normalized to mass concentration - typically, absorbance at 1% solution concentration at 1cm path length - then k becomes the specific absorption coefficient or dA/dc; sometimes written as [A1%1cm]. The units of dA/dc are g-1Lcm-1sup> .
If absorbance is normalized by molar concentration of the absorbing species, then k becomes the molar extinction coefficient or ε. The units of ε are mol-1 L cm-1.
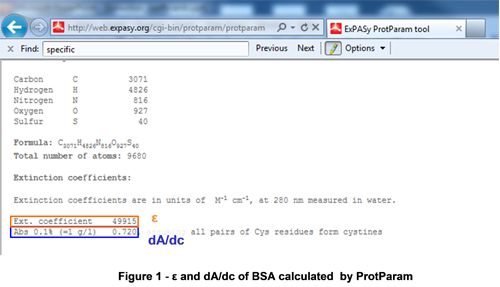
The molar extinction coefficient (ε) for proteins is often reported as the value obtained at 280 nm. This value is commonly calculated from the primary amino acid sequence in silico with computationaltools such as ProtParam (ExPASy http://web.expasy.org/cgi-bin; Figure 1).
dA/dc may be calculated for any compound if the molar extinction coefficient (ε) is known (Equation 2).
Equation 2
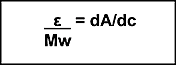
|
The parameter dA/dc is used in the OmniSEC software when UV is used as a concentration detector. The software will determine the concentration of the eluting sample from the absorbance at each data point. This value is then input into the Zimm equation to determine molecular weight.
dA/dc is also used when determining the composition of a conjugate species e.g. protein-detergent or protein-polymer complexes. The OmniSEC copolymer method uses RI and UV detectors to measure the concentration of both components in a conjugated sample. Light scattering is then used to measure the conjugate molecular weight.
|