Learn how ITC can be used to guide protein (RNA)/ligand crystallization and to resolve common issues with crystallization of complexes.
Eric Ennifar, Cyrielle Da Veiga, Dominique Burnouf, Joelle Mezher, Barbara Puffer-Enders and Philippe Dumas. Corresponding author: e.ennifar@unistra.fr Architecture et Réactivité de l’ARN, Institut de Biologie Moléculaire et Cellulaire, CNRS/Université de Strasbourg.
Crystallization of complexes (protein/protein, protein/nucleic acid, protein/ligand, nucleic acids/nucleic acid, nucleic acid/ligand), even on well-characterized biological systems, are frequently tedious and either time or sample consuming. The success rate of complex crystallization can be significantly improved if a proper preliminary characterization of the complex using biophysical methods is performed. It well established that Dynamic Light Scattering (DLS) is key to evaluate sample crystallizability (1). Likewise, DSC thermofluor-based optimization strategy have been developed to facilitate protein/ligand crystallization (2). Isothermal Titration Calorimetry (ITC) is the “gold standard” technique for investigating molecular interactions and we show here that it can be a valuable technique to improve crystallization of complexes. ITC is a true in-solution technique that directly provides, in one single experiment, the complete binding profile between two molecules: binding affinity (Ka), enthalpy and entropy changes (ΔH and ΔS) and stoichiometry (N) between two molecules are obtained very accurately (3-5). A major advantage of ITC over other similar biophysical assay approaches is that it is not restricted by macromolecule upper or lower size limit, there is no buffer restriction, and, most importantly for structural studies, it does not required any labeling. In addition, we have recently shown that modern ITC apparatus and new processing methods also allow obtaining a complete kinetic description on more diverse systems than usually thought, ranging from simple ligand binding to complex RNA folding (6).
The main limitation of ITC is the relatively large amount of sample required for an experiment. This should however not be a bottleneck for structural biologists involved into NMR or X-ray crystallography studies since sample requirement are quite similar to ITC analysis. Because the sample is not damaged during the ITC experiment, it can then be recovered and concentrated for subsequent crystallogenesis experiment. Finally, ITC can also be used to assess the proper folding of a nucleic acid or a protein used in a complex. We will present here some examples of such ITC-guided crystallization. All experiments were carried out on Microcal™ iTC200 (Malvern Instruments Ltd).
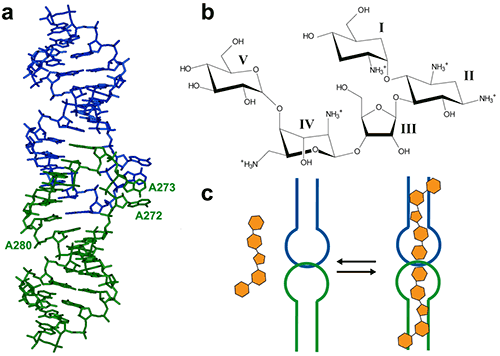
The HIV-1 genomic RNA Dimerization Initiation Site (DIS) is a highly conserved sequence with a stem-loop structure. The loop contains a self-complementary sequence that promotes dimerization of the viral genome by forming a loop-loop (kissing-loop) complex between two DIS hairpins. Crystal structures of the DIS loop-loop complex (7) revealed unexpected similarities with the bacterial 16 S ribosomal aminoacyl-tRNA site (A site), which is the target of aminoglycoside antibiotics (8). Consequently, it was shown that HIV-1 DIS RNA also binds aminoglycosides and can thus be an attractive target for the rationale design of new drugs targeting the HIV-1 genomic RNA (9). Here we will show how ITC can be used to guide co-crystallization of a complex made with the aminoglycoside Lividomycin and the HIV-1 DIS RNA kissing-loop complex.
|
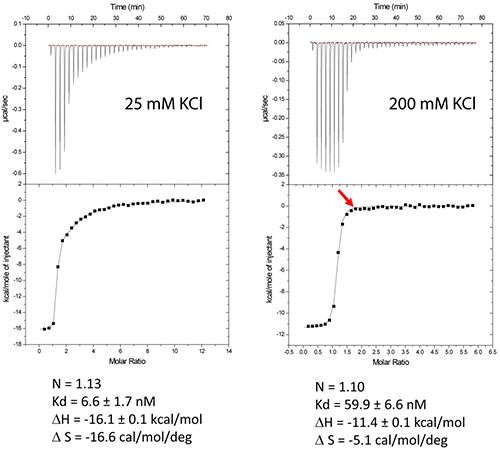
The DIS/lividomycin interaction provides an example of salt-dependent binding, as frequently observed with nucleic acids. As many nucleic acids ligands, aminoglycosides are positively charged and unspecific interactions due to electrostatic interactions are predominant at low salt concentration. However, excessive increase in salt concentration might prevent ligand binding due to electrostatic screening. One might therefore perform preliminary ITC experiments in various salt conditions in order to decrease unspecific binding while maintaining specific interactions. Experiments performed in a low-salt buffer (25 mM KCl, 2 mM MgCl2, 25 mM Na cacodylate pH 7.0) show two binding events and cannot be fitted with a single site model (Fig. 2A). Fitting with a two independent binding sites model shows a first very tight binding with a 1:1 stoichiometry, as expected (1 ligand for each stem-loop, 2 ligands for a kissing-loop complex), and a second binding site with an affinity in the low micromolar range that corresponds to unspecific interactions. This non-specific binding might therefore preclude formation of an homogenous RNA/ligand complex in crystallization conditions containing >100 mM concentration of the complex. However, in higher-salt conditions (200 mM KCl, 2 mM MgCl2, 25 mM Na cacodylate pH 7.0) only the specific and tight binding is observed (Fig. 2B). Such experimental conditions leading to a homogenous RNA/ligand complex should therefore be preferred in the search for crystallization conditions.
Misfolding is a frequent problem in structural studies of RNA. Here, the expected stoichiometry is obtained and these ITC experiments therefore also assess the proper folding of the RNA and, additionally, that the correct antibiotic concentration was used.
|
Another interesting perspective of ITC is the real-time monitoring of the stoichiometry in order to stop the experiment when the desired RNA/ligand ratio is reached (supposedly ≥ 1.0). For example in the DIS RNA/Lividomycin titration at 200 mM KCl, the experiment could be stopped at the 12th injection when RNA is saturated with the ligand (red arrow). This approach should be considered in situation where the stoichiometry can be easily estimated, i.e. when the slope of the resulting sigmoid is rather steep (Wieseman coefficient c ≥ 100) and when the off-rate constant (koff) is supposedly slow. In such a case, this approach prevents the presence of an excess of ligand in the sample and crystallization is performed with an optimal RNA/ligand complex ratio.
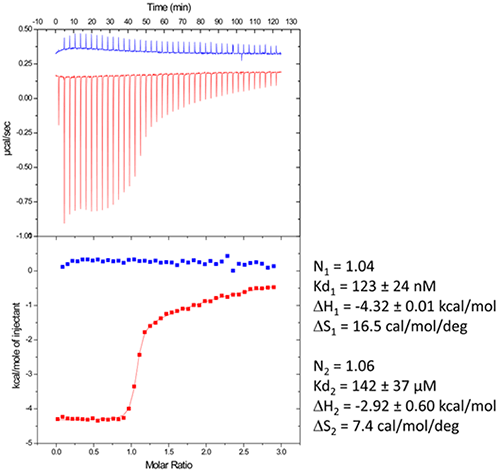
We use in the following a standard example with non-optimized salt concentration for the DIS kissing-loop RNA/Lividomycin interaction (ITC buffer made with 150 mM KCl, 5 mM MgCl2, 20 mM Sodium Cacodylate, pH 7.0.). The RNA sequence used is a chemically-synthesized 23-nucleotide long and to the HIV-1 subtype A DIS containing a bromouridine on the third position(7) in order to facilitate crystal structure determination using the multiwavelength anomalous dispersion (MAD) approach. The RNA sample at 120 µM concentration (in RNA strands) is prepared in the ITC buffer and placed in the ITC cell. The syringe is filled with Lividomycin 2.4 mM in the ITC buffer. ITC analysis is performed at 12°C in order to improve the complex formation (since the interaction is exothermic) and preserve the RNA from possible degradation.
|
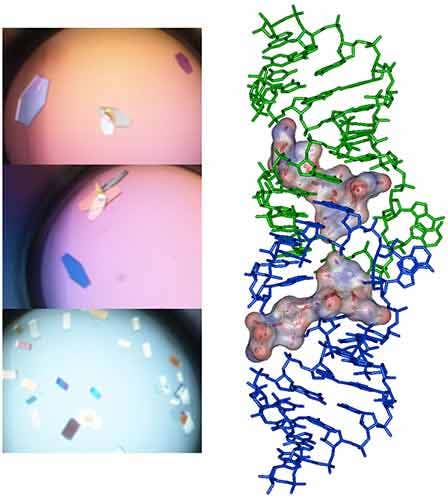
Following the ITC experiment, the complex formed in the ITC cell is then recovered and concentrated about three times (at 4°C to preserve the RNA/ligand interaction) to about 400 µM using microcentrators. This solution is then mixed (1/1 volume) in crystallization trays using commercial crystallization sparse matrix screens (containing 96 different crystallization conditions) and a crystallization robot. The crystallization reservoir is filled with the sparse matrix solution and trays are placed for incubation at 37°C. Large monocrystals suitable for diffraction (up to 1.6 Å resolution using a synchrotron X-ray source) and structure determination (Fig. 4) should grow after 16-24h incubation using Hampton Research® Natrix HT sparse matrix conditions (Fig. 4).
|
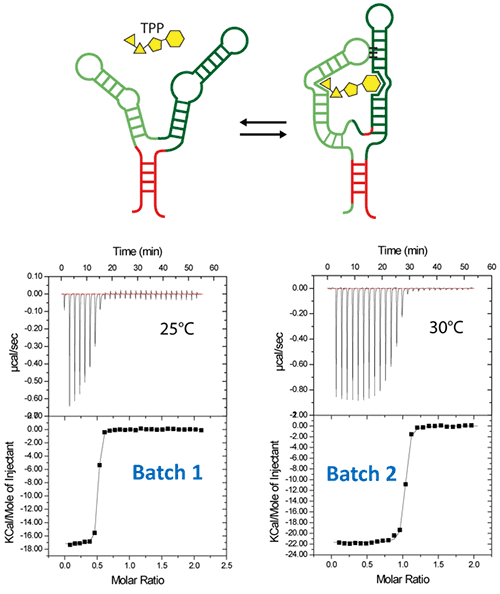
Riboswitches are untranslated regions of messenger RNA, which bind specific metabolites. Upon binding, a conformational switch is observed, thus regulating the expression of proteins involved in the biosynthesis of riboswitch substrates (12). The E.coli ThiC TPP riboswitch responds to the coenzyme thiamine pyrophosphate (TPP), an active form of vitamin B1 (13) (Fig. 5, top). We use in this example a 85-nucleotides synthetic RNA fragment corresponding to the Ecoli TPP aptamer domain(6). A major problem of such a rather large RNA with complex secondary and tertiary structures is to recover properly folded molecules after overproduction, purification, and in vitro renaturation process.
In a first step, the following refolding protocol was tried (RNA Batch 1): after purification, the RNA was heat-deturated, flash-cooled on ice and placed in the ITC buffer made with Na cacodylate pH 6.5 50mM, potassium acetate 100 mM, magnesium acetate 5 mM. Following RNA refolding, an ITC experiment is performed at 25°C: RNA at 30 µM is placed in the ITC cell and the syringe is loaded with TPP 500 µM. This ITC experiment shows that only about half of the RNA is able to bind the TPP ligand, which is a strong indicator of serious misfolding problem in the RNA (Fig. 5, middle, see “Batch 1”). The RNA/TPP complex formed in the ITC cell is then recovered, concentrated and used for crystallization trials as described above for the HIV-1 DIS kissing-loop RNA. However, only clear drops are observed from these crystallization assays (Fig. 5, bottom left), in agreement with the hypothesis that a major fraction of the RNA was misfolded.
In a second step, the RNA folding protocol was changed: after heat denaturation at 90°C for 3 min in H2O, the riboswitch was cooled to room temperature for 40 min. 10X ITC buffer was added to a final 1X concentration and the mix was further incubated for 40 min at room temperature. An ITC experiment performed on this second RNA batch (30 µM RNA, 500 µM TPP) show that this RNA refolding protocol lead to a fully active riboswitch since 100% of the RNA is able to bind the TPP ligand (Fig. 5, middle, see “Batch 2”). After concentration and setting up of crystallization drops, large monocrystals diffracting up to 5 Å resolution are obtained (Fig. 5, bottom right). This clearly ascertains the interest of using ITC as a quality control of the complex formation prior setting up crystallization trials.
|
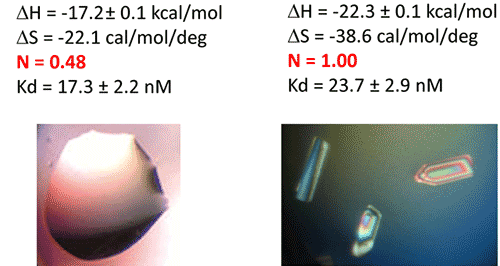

|
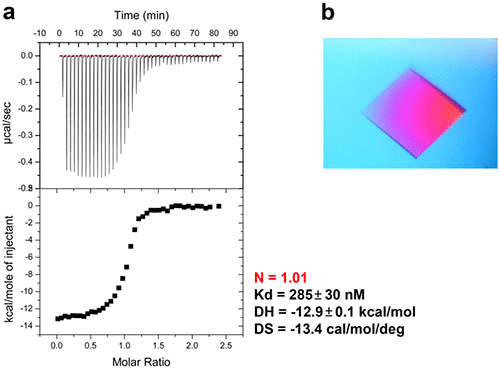
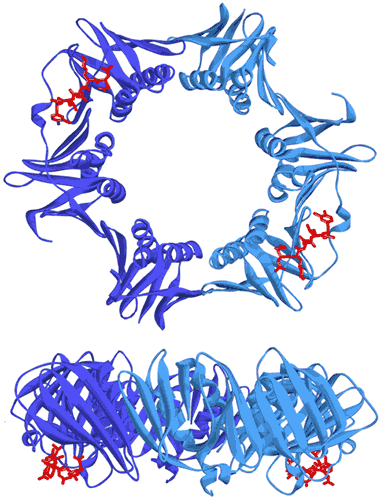
In bacteria, the DNA polymerase processivity factor, or β sliding clamp, is a homodimer that confers high processivity to DNA polymerases and other factors involved in DNA metabolism, through their interaction via a small conserved peptide into a hydrophobic pocket of the clamp (14). Short synthetic peptides have been developed in order to interfere with this interaction to prevent binding of the sliding clamp to the DNA polymerase, thus acting as antibiotics (15). The purpose of this study was to cocrystallize P.aeruginosa clamp bound to a synthetic peptide to get structural insights into this interaction and to aid the structure-based rationale design of new antibiotic peptides with a better affinity. Prior crystallization, an ITC experiment was performed by placing the β sliding clamp (35 µM in Hepes pH 7.4 10 mM, NaCl 150 mM, EDTA 3 mM) in the ITC cell and the peptide (300 µM) in the syringe (Fig. 6a). This complex was further used for crystallization trials after concentration to 47 mg/ml using a microcentrator. Crystallization was set by mixing one volume of protein/peptide complex with one volume of a reservoir solution made with 0.1 M sodium Mes pH 6.0 100 mM, CaCl2 100 mM and 20-30% (w/v) polyethylene glycol (PEG) 400. After several days of incubation at 20°C, large monocrystals diffracting up to 2.2 Å using a synchrotron radiation were obtain (Fig. 6b) and the X-ray structure was solved (Fig. 7).
|
|
The ITC technique can be implemented in the crystallization workflow of biological macromolecules complexes and can be used as a guide to improve the success rate of crystallization:
Sample requirement for structural studies are well-suited for ITC analysis
Assessment of the real active protein and/or ligand concentration (provided by the observed stoichiometry)
Real-time determination of an optimized protein/ligand ratio to be used for crystallization of the complex by monitoring the stoichiometry
Help in optimization of the complex formation (sample homogeneity, affinity, specificity)
Complete thermodynamic profile of the interaction is provided “for free”. One just needs to concentrate the sample after ITC analysis.
Ferre-D'Amare, A. R., and Burley, S. K. (1994) Use of dynamic light scattering to assess crystallizability of macromolecules and macromolecular assemblies, Structure 2, 357-359.
Dupeux, F., Rower, M., Seroul, G., Blot, D., and Marquez, J. A. (2011) A thermal stability assay can help to estimate the crystallization likelihood of biological samples, Acta Crystallogr D Biol Crystallogr 67, 915-919.
Ladbury, J. E., and Chowdhry, B. Z. (1996) Sensing the heat: the application of isothermal titration calorimetry to thermodynamic studies of biomolecular interactions, Chem Biol 3, 791-801.
Leavitt, S., and Freire, E. (2001) Direct measurement of protein binding energetics by isothermal titration calorimetry, Curr Opin Struct Biol 11, 560-566.
Privalov, P. L., and Dragan, A. I. (2007) Microcalorimetry of biological macromolecules, Biophys Chem 126, 16-24.
Burnouf, D., Ennifar, E., Guedich, S., Puffer, B., Hoffmann, G., Bec, G., Disdier, F., Baltzinger, M., and Dumas, P. (2012) kinITC: a new method for obtaining joint thermodynamic and kinetic data by isothermal titration calorimetry, J Am Chem Soc 134, 559-565.
Ennifar, E., Walter, P., Ehresmann, B., Ehresmann, C., and Dumas, P. (2001) Crystal structures of coaxially stacked kissing complexes of the HIV-1 RNA dimerization initiation site, Nat Struct Biol 8, 1064-1068.
Ennifar, E., Paillart, J. C., Marquet, R., Ehresmann, B., Ehresmann, C., Dumas, P., and Walter, P. (2003) HIV-1 RNA dimerization initiation site is structurally similar to the ribosomal A site and binds aminoglycoside antibiotics, J Biol Chem 278, 2723-2730.
Bernacchi, S., Freisz, S., Maechling, C., Spiess, B., Marquet, R., Dumas, P., and Ennifar, E. (2007) Aminoglycoside binding to the HIV-1 RNA dimerization initiation site: thermodynamics and effect on the kissing-loop to duplex conversion, Nucleic Acids Res.
Ennifar, E., and Dumas, P. (2006) Polymorphism of Bulged-out Residues in HIV-1 RNA DIS Kissing Complex and Structure Comparison with Solution Studies, J Mol Biol 356, 771-782.
Ennifar, E., Paillart, J. C., Bodlenner, A., Walter, P., Weibel, J. M., Aubertin, A. M., Pale, P., Dumas, P., and Marquet, R. (2006) Targeting the dimerization initiation site of HIV-1 RNA with aminoglycosides: from crystal to cell, Nucleic Acids Res 34, 2328-2339.
Winkler, W. C., and Breaker, R. R. (2005) Regulation of bacterial gene expression by riboswitches, Annu Rev Microbiol 59, 487-517.
Winkler, W., Nahvi, A., and Breaker, R. R. (2002) Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression, Nature 419, 952-956.
Burnouf, D. Y., Olieric, V., Wagner, J., Fujii, S., Reinbolt, J., Fuchs, R. P., and Dumas, P. (2004) Structural and biochemical analysis of sliding clamp/ligand interactions suggest a competition between replicative and translesion DNA polymerases, J Mol Biol 335, 1187-1197.
Wolff, P., Olieric, V., Briand, J. P., Chaloin, O., Dejaegere, A., Dumas, P., Ennifar, E., Guichard, G., Wagner, J., and Burnouf, D. Y. (2011) Structure-based design of short peptide ligands binding onto the E. coli processivity ring, J Med Chem 54, 4627-4637.
Wolff, P., Amal, I., Olieric, V., Chaloin, O., Gygli, G., Ennifar, E., Lorber, B., Guichard, G., Wagner, J., Dejaegere, A., and Burnouf, D. Y. (2014) Differential modes of peptide binding onto replicative sliding clamps from various bacterial origins, J Med Chem 57, 7565-7576