This application note primarily explores the applications of Isothermal Titration Calorimetry (ITC) in kinase inhibitor research, but touches on the role of Differential Scanning Calorimetry (DSC) as a quality control for the target protein in the discovery process.
The kinome of an organism is the set of protein kinases in its genome and these protein-modifying enzymes are potential targets in a number of therapeutic areas. Studies to characterize the human kinome, together with the explosion of available kinase crystal structures over recent years, have led to an increased focus on kinases as potential targets for pharmaceutical intervention.
Most kinase inhibitors that have been developed since the pharmaceutical industry became interested in kinases in the late 1980s target the ATP binding site of the enzyme. However, the development of the drug Gleevec™, a tyrosine-kinase inhibitor that induces a structural rearrangement of the target BCR-Abl kinase, served to reinvigorate research in this area. This has led to innovative ideas for tackling kinase inhibition, including binding outside of the ATP site and attempting to prevent kinase activation.
Detailed enzyme kinetic studies play an integral role in characterizing the mechanism of action of compounds based on novel ideas. One highly valuable technique is isothermal titration calorimetry (ITC) which provides a complete thermodynamic profile of the binding of a compound to the target protein. The data produced allow comparison of affinity measurements for compounds binding to different enzyme forms, for example free enzyme, enzyme-substrate complex, enzyme-product complex, active and non-active enzyme.
The inhibition of kinase signalling cascades is a proven therapeutic approach for the treatment of diseases in the areas of both oncology and inflammation. This application note focuses on the application of ITC to identifying the intermolecular complex of kinases that confers biological activity, knowing that ITC can provide information as to whether the presence of another ligand has an influence on that activity.
Isothermal titration calorimetry measures a number of characteristics of a binding interaction, including the affinity (KD), number of ligand binding sites (n), and the enthalpy of the binding reaction (∆H), in a single experiment. The technique is rapid, requires no fluorescent labelling and can be used with proteins that have no catalytic activity (which would preclude their study in enzyme kinetic assays).
ITC experiments usually involve titrating the test compound against the target protein at a constant temperature, with the ITC instrument measuring the heat released or taken up during the binding event. The technique can be used in a variety of ways during protein kinase-focused drug discovery.
These include:
characterization of protein constructs and preparations, not only in terms of correct affinity for model ligands, but also for correct stoichiometry, allowing estimation of the amount of functional protein, without the need for catalytic activity
evaluation of assays
identification of the intermolecular complex that confers biological activity, with ITC able to provide information on whether the presence of another ligand has an influence on the biological activity of a test compound, the focus of this application note.
Before mechanistic studies are carried out it is useful, if not essential, to undertake a quality control investigation of the target protein. This should take the form of verifying the identity, purity, concentration, functionality and stability of the protein.
Calorimetric methods can be applied in two of these important areas. ITC has been used to validate the functionality of target proteins by comparing the affinity and stoichiometry obtained on titration of a known ligand with literature values for a range of target proteins. The related technology of differential scanning calorimetry (DSC) has been used to verify that the protein has a melting temperature, Tm, significantly above the temperature of the experiment. Isolated kinase domains are often only partially stable, having Tm values around 40°C, (Figure 1). Applying techniques such as these to characterizing a target protein before embarking on detailed mechanistic studies can be time and cost-effective in the long term, helping avoid artefactual or misleading results due to poor protein quality. Low melting temperatures of kinase domains potentially indicate low stability and highlight the need for improved purification protocols, storage or assay conditions.
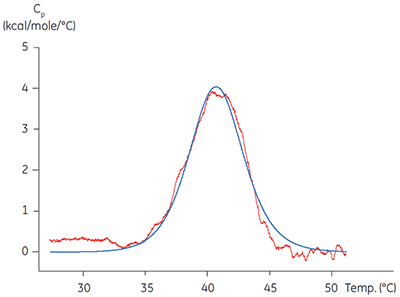
|
Information on whether the presence of another ligand influences the biological activity of a test compound is crucial to drug discovery. A second ligand may have: no effect on the activity of the test compound; may compete either directly or indirectly with the binding of the compound; or may actually be required for the test compound to exert its effect.
Understanding the mechanism of action of the test compound can be useful in interpreting or predicting cellular activity at substrate concentrations that are different from those used to measure IC50 values. It can also give insight into the relevance of 3D structures, which may be solved for different intermolecular complexes. Information on mechanism of action can also be used to devise subsequent assays, which can target a particular intermolecular complex. Kinase inhibitors may preferentially bind to, or induce non-active conformations of, the kinase protein. Comparing the binding affinity of a test compound for each of these forms is also valuable in deciding whether to pursue the development of compounds binding to the active or non-active form of the protein. Decisions of this type will affect the subsequent processes involved in kinase drug discovery.
As with all therapeutically important enzymes, kinases are not simply a single molecular target for compound intervention. During the catalytic cycle the kinase binds protein substrate, ATP, intermediates and products, (Figure 2). These different enzyme forms may also exist in many different conformations. Thus there are different forms of the enzyme to which compounds may be directed and to which different biochemical assays may be biased. The physiological concentration of ATP is around 2 mM, meaning that ATP is able to compete, often very effectively, with compounds binding at the ATP site. Therefore searching for compounds that bind before or after ATP (so-called non-competitive compounds) or those that bind only after ATP (so-called uncompetitive compounds) are both attractive approaches for kinase drug discovery. However, many historic kinase inhibitors target the ATP site of the enzyme and are expected to compete with ATP for binding to the enzyme. Other compounds may target allosteric sites, which may be expected not to show this competition with ATP. Characterizing the mechanism of action of compounds allows identification of whether the presence of ATP, or indeed protein substrate, increases, decreases or has no effect on the affinity of the test compound. Studies of this nature are valuable in understanding structure-activity relationships (SAR) at a molecular level and are key to the search for novel pharmacophores.
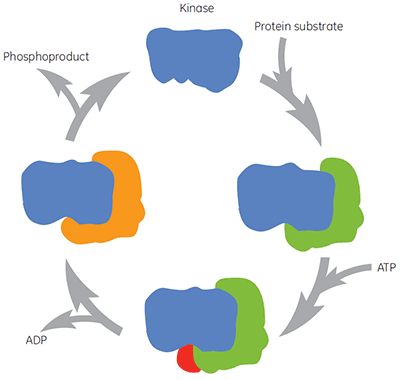
|
Enzyme kinetic assays usually are not configured to populate particular enzyme forms occurring along the reaction pathway. As a result it is sometimes difficult to directly obtain information on the relevant enzyme form for maximal affinity. ITC may overcome this limitation by measuring binding affinities to different, predetermined enzyme forms. Binding to free enzyme is the simplest approach, but ITC conditions can be arranged to probe other enzyme forms, such as `enzyme-protein substrate, enzyme-ATP, enzyme-ADP, or enzyme-phosphoproduct complexes, depending upon the mechanism of catalysis. The use of nonhydrolyzable ATP analogues can also be valuable in probing potential compound binding to the ternary complex of enzyme with both substrates.
Experiments to characterize the effect of ATP on the binding of a test compound were carried out for a protein kinase target. ITC titrations were carried out using the Malvern MicroCal™ VP-ITC for a test compound in the presence and absence of 100 µM ATP, representing approximately 60 × KD for ATP (Figure 3).
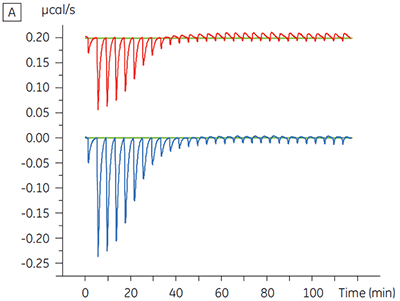
|
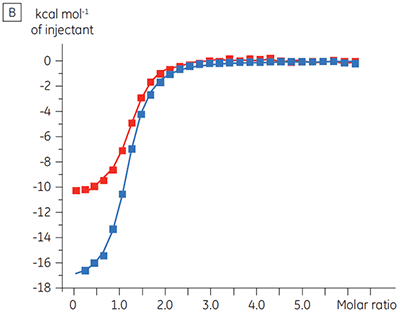
|
The ITC results show clearly that there is no change in affinity for the compound binding to the kinase when ATP is included with the protein in the cell. The enthalpy values indicate that although there is no effect on the affinity, there is a significant effect on the enthalpy of binding. These results therefore suggest that the compound is non-competitive with respect to ATP binding, and that there may be some change in binding mode in the presence of ATP. This highlights not only that ITC is useful in characterizing mechanistic details of compound binding, but also the dual probe nature of the technique is helpful in measuring not only affinity but also binding enthalpy. In a similar experiment, uncompetitive kinetics with respect to ATP were observed for a different test compound binding to the same protein kinase. The KD in the absence of ATP was > 50 µM (not measurable in the standard ITC run) whereas in the presence of ATP it was measured at 0.7 µM. (Figure 4).
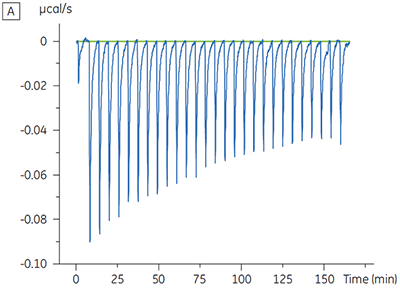
|
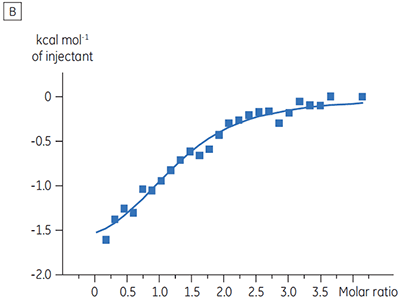
|
The advantage of being able to directly monitor binding to individual protein complexes was also shown in another example in which high affinity binding of a compound was suspected for a complex of two consecutive kinases in a signalling pathway. ITC allowed the binding of the compound to be studied to the upstream kinase alone and also to a complex of the upstream and downstream kinases, after first demonstrating that this complex did in fact form. A five-fold increase in affinity was demonstrated for the compound binding to the complex (Figure 5).
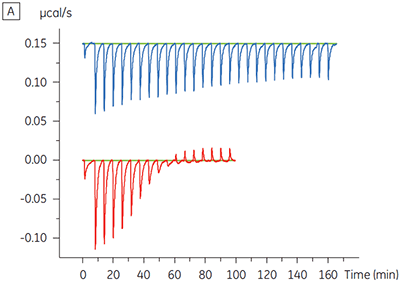
|
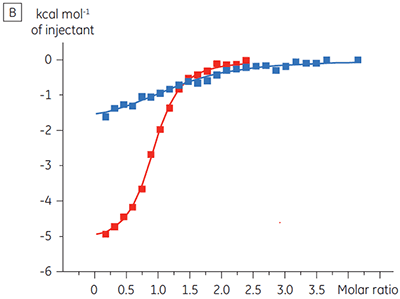
|
Calorimetric methods have proved to be valuable in the study of protein kinase inhibition. They facilitate quality control checks on target proteins as well as contributing to the dissection and understanding of inhibitor binding mechanisms.
The advent of high throughput and automated instruments, such as the Malvern MicroCal Auto-iTC200 and Malvern MicroCal VP-Capillary DSC systems, has added value to these applications. A continuing drive towards lower reagent consumption will ensure that calorimetric methods remain embedded in the process of rational drug design.
Geoff Holdgate, Principal Scientist, Global Compound Sciences, Lead Generation- Discovery Capabilities & Sciences, Astra Zeneca Pharmaceuticals, Mereside, Alderley Park, Macclesfield, SK10 4TG, UK