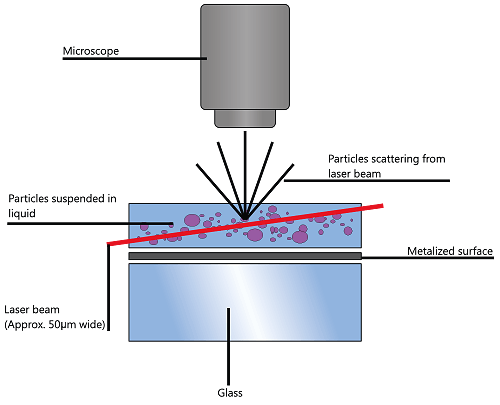
NTA utilizes the properties of both light scattering and Brownian motion in order to obtain the particle size distribution of samples in liquid suspension. A laser beam is passed through the sample chamber, and the particles in suspension in the path of this beam scatter light in such a manner that they can easily be visualized via a 20x magnification microscope onto which is mounted a camera. The camera, which operates at approximately 30 frames per second (fps), captures a video file of the particles moving under Brownian motion within the field of view of approximately 100 μm x 80 μm x 10 μm (Figure 1).
|
The movement of the particles is captured on a frame-by-frame basis. The proprietary NTA software simultaneously identifies and tracks the center of each of the observed particles, and determines the average distance moved by each particle in the x and y planes. This value allows the particle diffusion coefficient (Dt) to be determined from which, if the sample temperature T and solvent viscosity η are known, the sphere-equivalent hydrodynamic diameter, d, of the particles can be identified using the Stokes-Einstein equation (Equation 1).
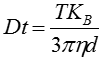
|
where KB is Boltzmann’s constant.
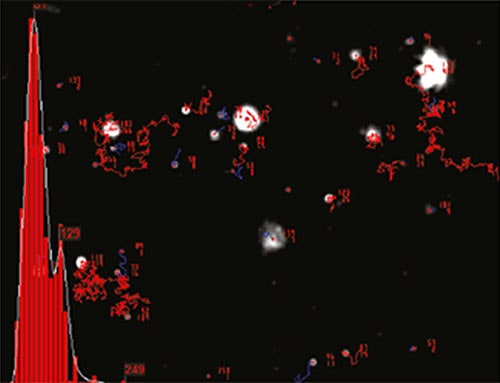
NTA is not an ensemble technique interrogating a very large number of particles, but rather each particle is sized individually, irrespective of the others. An example of the size distribution profile generated by NTA is shown in Figure 2.
|
In addition, the particles’ movement is measured within a fixed field of view (approximately 100 μm by 80 μm) illuminated by a beam approximately 10 μm in depth. These figures allow a scattering volume of the sample to be estimated; by measuring concentration of the particles within this field of view and extrapolating to a larger volume it is possible to achieve a concentration estimation in terms of particles per mL for any given size class or an overall total.
As NTA allows suspensions of nanoparticles to be visualized, sized and enumerated based on a particle-by-particle basis, its ability to determine the concentration and direct number frequency based particle size distribution profile means that virus preparations in particular can be studied in higher detail.
It is frequently the case in vaccine production and manufacture that the size of any particular virus or phage particle is known and of secondary importance to the estimation of virus particle concentration measurement and degree of aggregation. In this regard, the ability of NTA to determine virus concentration measurement through direct visualization, irrespective of virus infectivity, is of significant value.
Similarly, NTA’s ability to detect, size and, most importantly, measure concentration of all but the smallest viruses has led to numerous studies and assessment in this area, where it is considered to compare favorably with existing techniques.
Normally, the titre of phage and virus particles is established by plaque assay or, in the case of animal cell viruses, by cell based bioassay. In these assay systems, infective virus particles are grown in confluent cell layers to produce plaques (zones of destroyed cells) which can be detected. While a direct count of individual infective virus particles can be obtained, non-infective virus particles do not produce plaques and therefore cannot be detected. In addition, aggregates containing many virus particles will produce only single plaques. Often, the manufacturer needs to know the number of virus particles in the preparation, whether infective or not, and the degree, if any, to which the preparation is undergoing aggregation (an early indicator of limited product shelf life).
Accordingly, NTA has been assessed for this purpose. Early work established the technique’s potential for fast and low cost analysis of small volume samples of high polydispersity but recognized the then early stage of development and limited data on which to base estimates of reproducibility and accuracy (Moser, 2008).
Subsequent studies have furnished much more detailed information. In testing the ability of NTA to determine number concentration of both artificial and natural nanoparticles (such as adenoviruses), Du and his colleagues concluded that NTA had improved accuracy compared to mathematical calculation and spectrophotometer methods (Du et al. (2010)), although the concentration range over which NTA was applicable was a potential limitation. Nevertheless, it was shown that NTA could have significant applications in a number of fields ranging from nanoparticle synthesis, aggregation, and drug delivery to vaccine production. The same group also used NTA to study the adhesion of polystyrene and virus particles (Kendall et al. (2010)).
More recently, Kramberger et al. (2012) have evaluated NTA for total virus particle determination by testing its ability to quantify latex particles, adenovirus 5 and influenza virus over several consecutive days by using known concentrations of the subject particles. NTA analysis was also used to quantify chromatographic fractions of adenovirus and influenza virus after purification on a CIM monolithic column. NTA results were compared and evaluated against hemagglutination (HA) and end point dilution assay, determining total and infection virus particle number, respectively. The results demonstrated that Nanoparticle Tracking Analysis is a method for fast estimation of virus concentration in different samples. In addition, it can provide a better insight into the sample status, regarding the level of virus aggregation.
NTA has also been compared to conventional plaque assay (PA) and quantitative polymerase chain reaction (qPCR) for the detection and analysis of 3 phage types in a comprehensive study (Anderson et al. (2011)). Here, it was concluded that, while NTA only operates best in a relatively clear medium over an optimum concentration range and, compared to conventional PA methods, is more capital expensive, the technique generated results within an impressive ≤5 min timeframe, which was significantly faster than PA and the qPCR method (18-24 hours and 3-4 hours, respectively) and that its performance does not require any additional reagents. The authors suggested that, once optimized for phage, it would be likely that the NTA-based method will be reproducible among various laboratories, with accuracy comparable to PA performed by various investigators. However, it would be significantly faster and may be very useful for future basic and applied research with bacteriophage.
Similarly, Filipe et al. (2011) compared NTA to qPCR as a means by which a greater understanding of viral preparations can be gained in support of traditional and frequently limited techniques (Filipe et al. (2011)). They emphasized that while the qPCR technique is advantageous, in that it is highly specific and can be used to quantify DNA containing viruses in a harvest material, it cannot distinguish the amount of aggregation within a sample preparation. In addition, the technique measures only viruses that actually contain DNA or RNA (filled capsids) while many viruses formed in vaccine or gene therapy production contain no RNA or DNA (empty capsids) and thus cannot be measured using qPCR. The fact that NTA visualizes, sizes and measures concentration of all virus particles and, importantly, aggregates thereof allows the user to calculate the ratio of filled and empty capsids (despite varying levels of aggregates in a given sample). They concluded that the greatest value of NTA in this field is in making it possible to quantify, visualize and size not only viruses, but also aggregates. Finally they recognized the potential of NTA in its ability to measure fluorescently labelled virus particles thus extending the usefulness of this technique into harvest materials, for which it is essential to discriminate viruses from background host-cell debris and proteins.
In his study of the use of viral quantitative capillary electrophoresis (Viral qCE) for measuring concentration of intact viruses, Mironov used NTA to verify the quantitation of intact oncolytic Vaccinia virus particles by Viral qCE after calibrating the instrument with 100, 200 and 400 µm latex beads (Mironov et al. (2011)). In another study involving the detection of oncolytic viruses using small molecule single stranded RNA/DNA oligonucleotide aptamers, Chechik and Berezovski (2011) used NTA to evaluate the binding of DNA aptamers to Vaccinia virus and vesicular stomatitis (VSV). They showed that aptamers conjugated with beads caused the aggregation of virus particles and the intensity of light scattered by the aggregates was directly proportional to the affinity of aptamer pools to viruses. They concluded that NTA brings a valuable tool for all further developments of biomolecular binding analyses and aptamer selection.
Vesina et al. (2012) also obtained particle size distribution on their plant-derived virus-like particles in their development of methodology for their preparation. Trifonova et al. (2012a and 2012b), on the other hand, used structurally modified plant viruses as nanovaccines using NTA to assist in their characterization, while Azizi et al. (2012) have most recently used NTA to discriminate between variations in binding of the fluorescent label YOYO-1 to viral and ribosomal RNA differential binding in their development of a quantitative capillary electrophoresis technique for concentration measurement and quality control of RNA virus.
The use of lentiviral vectors for gene therapy necessitates the generation of functional vectors using fast, non-laborious and cost-effective strategies. Eleni et al. (2013) have described an improved protocol in which vector stocks are prepared by transient transfection using standard cell culture media and are then concentrated by ultrafiltration, resulting in functional vector titres of up to 6 x 109 transducing units per milliliter (TU/mL) without the involvement of any purification step. They determined the viral functional titre by employing flow cytometry and evaluated the actual viral particle size and concentration in real time by using NTA. Searing et al. (2013) have also recently reported characterizing lentiviral vector particles by NTA-determined size and composition.
The potato virus X (PVX) virion can be reconstituted in vitro from the virus coat protein (CP) and RNA with heterologous RNAs capable of being used as well. In a recent study, Petrova et al. (2013) showed the structure and properties of cognate and heterologous viral ribonucleoproteins (vRNPs) to be similar to those of native virions, using NTA to analyze an incubation mixture which was diluted 10,000 times in 10 mM Tris-HCl buffer, pH 7.5, to a protein concentration of 0.5 ng/mL.
As one of the vectors of choice for gene delivery and expression of foreign proteins in gene therapy and vaccination purposes, recombinant adenoviruses are highly efficient at gene transfer for a broad spectrum of cell types and species. Silva et al. (2013) have used NTA in the development of new scalable and reproducible production processes of adenoviral vectors (human and canine) using stirred tank bioreactors.
The use of NTA in vaccine research has been reviewed recently by several research groups.
In a study aimed at the development of large scale production protocols of virus-like particles (VLPs) which offer great promise as candidates for new vaccine strategies, Cervera et al. (2013a) described the generation of HIV-1 Gag VLPs by transient transfection of HEK 293 suspension cell cultures using an optimized animal-derived component free medium, as compared to the more conventional use of the baculovirus expression system. They used NTA to show that the maximum cell density attained using the optimized Freestyle culture medium was 5.4 x 106 cells/mL in batch mode, almost double of that observed using the unsupplemented medium (2.9 x 106 cells/mL). Cervera et al. (2013b) subsequently extended this work to demonstrate that the human embryonic kidney 293 (HEK 293) is the preferred host system due to the many industrially relevant features this cell line offers including high transfectability, ability to grow in suspension, ability to grow to high cellular densities and adaptation to serum-free culture conditions. The great majority of Gag-GFP recovered from cell culture supernatants was shown to be correctly assembled into VLPs of the expected size and morphology as demonstrated by NTA. The scalability of this strategy was demonstrated using a 1L WAVE bioreactor and the quantity and quality of Gag VLPs generated using the described platform was considered suitable for pre-clinical studies in mice. The same group has shown that, as an alternative to time-consuming ELISA based assays, the development and validation of a new fluorescence-based quantitation assay for Gag VLP titres were in good agreement with those obtained using TEM and NTA (Gutiérrez-Granados et al. (2013a)). They concluded that this simple, rapid and cost-effective quantitation assay should facilitate development and optimization of bioprocessing strategies for Gag-based VLPs. Gutiérrez-Granados et al. (2013b) have most recently reported on the characterization and quantitation of fluorescent Gag virus-like particles. Using a Gag-GFP fusion construct, the viral particle titres estimated were compared with those obtained by p24 ELISA (Innotest®, Innogenetics, Belgium), densitometry, TEM and NTA. Upon transfection, Gag-GFP was expressed in the cytoplasm of the producer cells and accumulated in the vicinity of the plasma membrane where the budding process takes place.
Mather (2013) has described the work of the Lomonosov Moscow State University (MSU; Moscow, Russia) who used NTA technology for vaccine characterization and standardization, as the technology enabled real-time measurement of virus particles in liquid without contamination or particle aggregation (clustering). The results enabled the research team to analyze and control the size, state of aggregation, and concentration of artificial plant virus particles and small spherical plant and animal viruses. It also allowed them to see the formation of immunogenic complexes (candidate vaccines) by using the indirect immunofluorescence or immunogold staining methods
Similarly, Patois et al. (2013) recently reviewed the applications of NTA for the characterization of biopharmaceuticals and procedures were proposed for the determination of size distributions. Size distributions of two subunit seasonal influenza vaccines (a) Influvac (Abbott) and (b) Agrippal (Novartis) were measured within 60 seconds using NTA. The review also discussed techniques such as fluorescence microscopy with Nile Red staining, micro-flow imaging, 90° light scattering, and flow field-flow fractionation (FFF) and concluded that NTA is a suitable technique for a wide range of applications in the field of biopharmaceuticals.
In the development of virus-like particles for the study of virus-host interaction of hepatitis E virus (HEV) Kapur et al. (2011) used TEM and NTA to show near uniform particles of approximately 30–35 nm in diameter for pORF2 VLPs and 60–100 nm for reporter-linked VLPs, while Hughson et al. (2010) described inactivation studies on Japanese Encephalitis Virus vaccine.
Aimed at the development of anti-cancer vaccines, work undertaken by Mayer-Sonnenfeld et al. (2013) compared protein content of chaperone-rich cell lysate (CRCL) anti-cancer vaccines prepared from human tumors of different histological origins to evaluate the uniformity of their protein content. Protein samples were separated and identified by SDS-PAGE and slices cut from gels for protease digestion followed by mass spectrometry analysis, the content assessed by gene ontogeny and networking programmatic computation. CRCL preparations were also analyzed by NTA and TEM.
Mann et al. (2013) showed that pulmonary delivery of DNA vaccine constructs using deacylated PEI elicited immune responses and protected against viral challenge infection. They used NTA to verify polyplex and lipoplex formations.
Cayatte et al. (2013) have been working on the development of a vaccine against human cytomegalovirus (HCMV), a betaherpesvirus, which can cause severe disease in immunosuppressed patients following congenital infection. Proposing that dense bodies (DB) are complex, non-infectious particles produced by HCMV infected cells and may represent a vaccine option, they explored characterization and defined DB production methods followed by systematic evaluation of neutralization and cell mediated immune responses to the DB material in Balb/c mice. They purified DB from Towne infected cultures treated with viral terminase inhibitor 2-bromo-5,6-dichloro-1-beta-d-ribofuranosyl benzimidazole riboside (BDCRB) which were characterized by NTA, 2D fluorescence difference gel electrophoresis (2D-DIGE), immunoblotting, quantitative ELISA and other methods.
Recently, Liu et al. (2013), working on influenza virosomes supplemented with GPI-0100 adjuvant, have evaluated, using NTA, the adjuvant effect of GPI-0100 on a virosomal influenza vaccine formulation. In contrast to influenza subunit vaccine adjuvanted with GPI-0100, highly immunogenic virosomal vaccine supplemented with the same dose of GPI-0100 provided full protection of mice against infection at extremely low hemagglutinin antigen doses, and allows the use of very low antigen doses without compromising the protective potential of the vaccine.
Patil et al. (2013) have recently evaluated monophosphoryl lipid A (MPLA) as an adjuvant for pulmonary delivered whole inactivated virus influenza vaccine (WIV). Using NTA to successfully size WIV derived from A/PR/8/H1N1 before and after addition of MPLA, these studies suggested that the mucosal and systemic immune responses to pulmonary delivered influenza vaccines can be significantly enhanced by using MPLA as adjuvant and that MPLA-adjuvanted SFD vaccine was particularly effective, implying that delivery of adjuvanted vaccine powder to the lungs can be an attractive way of immunization against influenza.
In their systematic experimental exploration into how different bacteriophage (phage) traits may influence the formation of a plaque, Gallet et al. (2011) constructed a series of isogenic λ phages that differ in their adsorption rate, lysis timing or morphology to determine the effects these changes have on three plaque properties: size, progeny productivity, and phage concentration within plaques. NTA was successfully used to determine phage concentration to conclude that a more realistic theoretical approach to plaque formation is needed in order to capture the complex interaction between phage and its bacterium host in a spatially restricted environment.
Phage were also the subject of another study by Zhou et al. (2012) in which NTA was used to confirm the high purity and dispersion of the 72nm fd Y21Mphage preparation used. Similarly, Carter et al. (2012) demonstrated that a commercially available bacteriophage cocktail (EcoShield™) significantly reduces Escherichia coli O157:H7 contamination of lettuce and beef, but did not protect against recontamination.
In the past decade, spherical and rod-like viruses had been used for the design and synthesis of a new kind of nanomaterials with unique chemical positioning, shape, and dimensions in the nanosize regime. Uniquely, NTA allows the average intensity of a particle’s scattering to be measured at the same time as its size is being determined by its dynamic behavior. This allows particles of differing refractive index, but same size, to be discriminated by plotting these two parameters as a function of each other. This capability has been exploited in experiments where virus particles, being of highly defined structure and uniform in size, have been used as templates for the simple production of highly monodisperse metallized particles by an electrodeless deposition metallization process.
Aljabali et al. (2011a) demonstrated chemically-coupled-peptide-promoted virus nanoparticle templated mineralization using Cowpea mosaic virus (CPMV). The cationic polyelectrolyte, poly(allylamine) hydrochloride (PAH), is electrostatically bound to the external surface of the virus capsid; the polyelectrolyte promotes the adsorption of anionic gold complexes, which are then easily reduced, under mild conditions, to form a metallic gold coating. As expected, the templated gold nanoparticles can be further modified with thiol reagents (Aljabali et al. (2011b)).This work followed earlier studies in which they used NTA to demonstrate mineralization of the virus. For each of the mineralized virus particles there was a significant increase in the relative refractive index compared with wild-type and with peptide-CPMV conjugates. The analysis was also considered consistent with the mineralized particles being monodisperse as indicated by the particle size distribution; although for ZnS-CPMV there was a little non-specific aggregation under the measurement conditions employed (Aljabali et al. (2010)). NTA is unique in its ability to measure both the size and differences in light scattering properties of particles on an individual basis.
The plant viral nanoparticle tobacco mosaic virus (TMV) has been used for the production of a variety of metals including cobalt, nickel, iron, platinum, cobalt–platinum and nickel–iron and gold nanowires, in which successful deposition of highly refractive metal layers to the surface of the viruses were seen as an increase in scattered intensity without a significant change in particle size which would have indicated aggregation (Bromley et al. (2008)). Other studies have prepared virus templated nanoparticles of silica (Steinmetz et al. (2009)) and iron-platinum (Shah et al. (2009)).
To compensate for the low sensitivity of magnetic resonance imaging (MRI), nanoparticles have been developed to deliver high payloads of contrast agents to sites of disease. Bruckman et al. (2013) recently reported the development of supramolecular MRI contrast agents using rod-shaped TMV nanoparticles (300 × 18 nm) that were loaded with up to 3500 or 2000 chelated paramagnetic gadolinium(III) ions selectively at the interior or exterior surface, respectively. Dynamic Light Scattering (DLS) and NTA were used to show that interior-labelled TMV rods can undergo thermal transition to form 170 nm sized spherical nanoparticles containing 25 000 gadolinium chelates and a per-particle relaxivity of almost 400 000 mM−1 s−1 (15.2 mM−1 s−1 per gadolinium), thus laying the foundation for the use of TMV as a contrast agent for MRI. Other studies on biotemplating rod-like viruses for the synthesis of copper nanorods and nanowires have been reported recently (Zhou et al. (2012)) in which was demonstrated the controlled synthesis of copper nanorods and nanowires by electroless deposition of copper on three types of palaydium-activated rod-like viruses. The synthesis conditions described in the work were considered scalable and amenable for biological templates and the synthesized structures preserve the dimensions and shape of the rod-like viruses utilized during the study. The work opened the possibility of generating a variety of nanorods and nanowires of different lengths ranging from 300 nm to micron sizes with potential in nanoelectronics, sensing, and cancer therapy. Finally, Trifonova et al. (2012a and b) have investigated the antigenic properties of complexes based on structurally modified plant viruses.
Aljabali AA, Barclay JE, Lomonossoff GP and Evans DJ (2010) Virus templated metallic nanoparticles, Nanoscale, 2010, Advance Article, DOI: 10.1039/C0NR00525H , Paper
Aljabali AA, Lomonossoff GP and Evans DJ (2011a) CPMV-polyelectrolyte-templated gold nanoparticles, Biomacromolecules, Just Accepted Manuscript DOI: 10.1021/bm200499v, Publication Date (Web): June 9, 2011
Aljabali AA, Shah SN, Evans-Gowing R, Lomonossoff GP and Evans DJ (2011b) Chemically-coupled-peptide-promoted virus nanoparticle templated mineralization, Integrative Biology, 2011, 3, 119-125
Anderson, B., Rashid MH, Carter C, Pasternack G, Rajanna C, Revazishvili T, Dean T, Senecal A and Sulakvelidze A (2011) Enumeration of bacteriophage particles: Comparative analysis of the traditional plaque assay and real-time QPCR- and NanoSight-based assays, Bacteriophage, Volume 1, Issue 2 March/April 2011
Azizi A, Mironov GG, Muharemagic D, Wehbe M, Bell JC and Berezovski MV (2012) Viral Quantitative Capillary Electrophoresis for Counting and Quality Control of RNA Virus, Anal. Chem., Just Accepted Manuscript, DOI: 10.1021/ac302525y, Publication Date (Web): October 9, 2012
Bromley KM, Patil AJ, Perriman AW, Stubbs G and Mann S (2008) Preparation of high quality nanowires by tobacco mosaic virus templating of gold nanoparticles, J. Materials. Chem, 18, 4796 – 4801.
Bruckman MA., Hern S, Jiang K, Flask CA, Yu X and Steinmetz NF (2013) Tobacco mosaic virus rods and spheres as supramolecular high-relaxivity MRI contrast agents, J. Mater. Chem. B, 2013, Advance Article, doi: 10.1039/c3tb00461a
Carr B, Hole P, Malloy A, Nelson P, Wright M. and Smith J (2009) “Applications of nanoparticle tracking analysis in nanoparticle research - a mini-review”, European Journal of Parenteral & Pharmaceutical Sciences 2009; 14(2): 45-50
Carter CD, Parks A, Abuladze T, Li M, Woolston J, Magnone J, Senecal A, Kropinski AM and Sulakvelidze A (2012) Bacteriophage cocktail significantly reduces Escherichia coli O157:H7 contamination of lettuce and beef, but does not protect against recontamination, Bacteriophage 2:3, 1–8; July/August/September 2012; c 2012 Landes Bioscience
Cayatte C, Schneider-Ohrum K, Wang Z, Irrinki A, Nguyen N, Lu J, Nelson C, Serva E, Gemmell L, Citkowicz A, Liu Y, Hayes G, Woo J, Van Nest G, Jin H, Duke G and McCormick AL (2013) Cytomegalovirus vaccine strain Towne-derived dense bodies induce broad cellular immune responses and neutralizing antibodies that prevent infection of fibroblasts and epithelial cells. Journal of Virology, Published ahead of print 7 August 2013, doi: 10.1128/JVI.01554-13
Cervera L, Gutiérrez S, Martínez M, Blanco J, Gòdia F, Segura MM (2013a) Generation of HIV-1 Gag VLPs by transient transfection of HEK 293 suspension cell cultures using an optimized animal-derived component free medium Journal of Biotechnology, http://dx.doi.org/10.1016/j.jbiotec.2013.05.001
Cervera L, Gutiérrez-Granados S, Gòdia F, Segura MM (2013b) HIV-1 VLP production platform based on transient transfection of HEK 293 suspension cultures in bioreactors. AIDS2013, http://epostersonline.s3.amazonaws.com/aids2013/aids2013.1af0d7b.NORMAL.pdf
Chechik A. and Berezovski M.(2011) Selection of Aptamers Against Oncolytic Viruses and Their Binding Evaluation by Nanoparticle Tracking, 94th Canadian Chemistry Conference and Exhibition, June 5–9, 2011, Palais des congrès de Montréal, Quebec, Canada,
Du S, Kendall K, Morris S, Sweet C (2010) Measuring number-concentrations of nanoparticles and viruses in liquids on-line, Journal of Chemical Technology & Biotechnology, Published Online: 19 May 2010, DOI: 10.1002/jctb.2459
Eleni P, Kontostathi G, Drakopoulou E, Georgomanoli M, Stamateris E, Vougas K, Vlahou A, Maloy A, Ware M, Anagnou N P (2013) Characterization and comparative performance of lentiviral vector preparations concentrated by either one-step ultrafiltration or ultracentrifugation, Virus Research, Available online 11 April 2013, http://dx.doi.org/10.1016/j.virusres.2013.03.015
Filipe V, Jiskoot W, Hawe A (2011) Understanding Virus Preparations Using Nanoscale Particle Characterization, BioProcess International, Vol. 9, No. 2, February 2011, pp. 44–51
Gallet R, Kannoly S and Wang I-N (2011), Effects of bacteriophage traits on plaque formation, BMC Microbiology 2011, 11:181 doi:10.1186/1471-2180-11-181
Gutiérrez-Granados S, Cervera L, Gòdia F, Carrillo J, Segura M M (2013a) Development and validation of a quantitation assay for fluorescently tagged HIV-1 virus-like particles, Journal of Virological Methods, http://dx.doi.org/10.1016/j.jviromet.2013.05.010
Gutiérrez-Granados S, Cervera L, Gòdia F, Segura MM (2013b) Characterization and quantitation of fluorescent Gag virus-like particles, BMC Proceedings 2013, 7(Suppl 6):P62 doi:10.1186/1753-6561-7-S6-P62
Hughson MD., Venables D, Queen K, Titchener-Hooker N and Mukhopadhyay TK. (2010), Inactivation studies on Japanese Encephalitis Virus Vaccine using Design of Experiments, in "Vaccine Technology III", (John G. Auniņš, Barry C. Buckland, Kathrin U. Jansen, Paula Marques Alves, Eds), ECI Symposium Series, Volume P13 http://services.bepress.com/eci/vaccine_iii/31
Kapur N, Thakral D, Durgapal H and Panda SK (2011) Hepatitis E virus enters liver cells through receptor-dependent clathrin-mediated endocytosis. Journal of Viral Hepatitis. doi: 10.1111/j.1365-2893.2011.01559.x
Kendall K , Du S, Morris S and Sweet C (2010) 'Virus Concentration and Adhesion Measured by Laser Tracking', The Journal of Adhesion, 86:10, 1029 – 1040
Kramberger P, Ciringer M, Strancar A, Peterka M (2012) Evaluation of nanoparticle tracking analysis for total virus particle determination, Virology Journal 2012, 9:265 doi:10.1186/1743-422X-9-265
Liu H, de Vries-Idema J, ter Veer W, Wilschut J, Huckriede A (2013) Influenza virosomes supplemented with GPI-0100 adjuvant: a potent vaccine formulation for antigen dose sparing, Medical Microbiology and Immunology, September 2013, DOI 10.1007/s00430-013-0313-2
Mach H and Arvinte T (2011) Addressing new analytical challenges in protein formulation development, European Journal of Pharmaceutics and Biopharmaceutics, Article in Press doi:10.1016/j.ejpb.2011.03.001 Mach and Arvinte (2011) have discussed NTA among other techniques for the analysis of sub-visible particles in protein therapeutics and formulation development
Maddux NR, Joshi SB, Volkin DB, Ralston JP and Middaugh CR (2011) Multidimensional methods for the formulation of biopharmaceuticals and vaccines. Journal of Pharmaceutical Sciences, 100: n/a. DOI: 10.1002/jps.22618
Mann JFS, McKay PF, Arokiasamy S, Patelb RK, Kleina K, Shattock RJ (2013) Pulmonary delivery of DNA vaccine constructs using deacylated PEI elicits immune responses and protects against viral challenge infection, Journal of Controlled Release, Available online 14 June 2013, http://dx.doi.org/10.1016/j.jconrel.2013.06.004
Mayer-Sonnenfeld T, Har-Noy Ml, Lillehei KO, and Graner MW (2013) Proteomic analyses of different human tumour-derived chaperone-rich cell lysate (CRCL) anti-cancer vaccines reveal antigen content and strong similarities amongst the vaccines along with a basis for CRCL's unique structure: CRCL vaccine proteome leads to unique structure International Journal of Hyperthermia, Ahead of Print : Pages 1-8, (doi: 10.3109/02656736.2013.796529)
Mather L (2013) Nanoparticle tracking analysis characterizes vaccines, BioOptics World, 01/29/2013, http://www.bioopticsworld.com/articles/2013/01/nanoparticle-tracking-analysis-characterizes-vaccines.html
Moser M (2008) Emerging analytical techniques to characterize vaccines, Proc. Intl. Conf. Vaccines Europe, Brussels, December 2008.
Mironov G G, Chechik AV, Ozer R , Bell JC , and Berezovski MV (2011) Viral Quantitative Capillary Electrophoresis (Viral qCE) for Counting Intact Viruses, Anal. Chem., Just Accepted, Publication Date (Web): May 21, 2011 (Article), DOI: 10.1021/ac201006u
NanoSight Limited (2013) www.nanosight.com
Patil H, Murugappan S, ter Veer W, Meijerhof T, de Haan A, Frijlink HW, Wilschut J, Hinrichs WLJ, Huckriede A (2013) Evaluation of monophosphoryl lipid A as adjuvant for pulmonary delivered influenza vaccine, Journal of Controlled Release, Available online 21 November 2013, http://dx.doi.org/10.1016/j.jconrel.2013.11.013
Patois E, Capelle MAH, Palais C, Gurny R, Arvinte T (2013) Evaluation of nanoparticle tracking analysis (NTA) in the characterization of therapeutic antibodies and seasonal influenza vaccines: pros and cons, J. Drug Del. Sci. Tech., 22 (5) 427-433 2012
Petrova EK, Nikitin NN, Protopopova AD, Arkhipenko MV, Yaminsky IV, Karpova OV, Atabekov JG (2013) The role of the 5′-cap structure in viral ribonucleoproteins assembly from potato virus X coat protein and RNAs, Biochimie, 2013, http://dx.doi.org/10.1016/j.biochi.2013.09.004
Shah SN, Steinmetz NF, Aljabali AAA, Lomonossoff GP and Evans DJ (2009) Environmentally benign synthesis of virus-templated, monodisperse, ironplatinum nanoparticles. Dalton Transactions, Cambridge (40):8479-80
Searing I, Bannister R, Jones P, Lad Y, Mitrophanous K (2013) Characterising lentiviral vector particles by size and composition, 10th Annual Conference of the British Society of Gene and Cell Therapy, Royal Holloway, University of London, 17th-19th April, 2013
Silva AC, Fernandes P, Sousa MFQ, Alves PM (2013) Scalable Production of Adenovirus Vectors, Adenovirus; Methods in Molecular Biology Volume 1089, 2014, pp 175-196
Steinmetz N, Shah SN, Barclay JE, Rallapalli G, Lomonossoff GP and Evans DJ (2009) Virus-Templated Silica Nanoparticles, Small, Volume 5, Issue 7, pages 813–816.
Trifonova EA, Nikitin NA, Karpova OV (2012a) The investigation in liquid of the antigenic properties of complexes, based on structurally modified plant viruses, 6th international conference “Progress and trends in bionanoscopy”, Moscow, 18-20 June 2012
Trifonova EA, Nikitin NA, Lazareva EA, Karpova OV, Atabekov IG. (2012b) The investigation of nanovaccines assembled in vitro with various techniques of bionanoscopy, 6th international conference “Progress and trends in bionanoscopy”, Moscow, 18-20 June 2012
Vezina L-P; Couture M; Paquet D; Dargis M; d'aoust M-A (2012) Method Of Preparing Plant-Derived VLPS, United States Patent Application 20120178149
Zhou JC, Soto CM, Chen MS, Bruckman MA (2012) Preparation of fd Y21M phage nanoparticles, www.jnanobiotechnology.com/content/supplementary/1477-3155-10-18-s1.doc