Protein interactions are strongly influenced by electrostatic forces. While hydrogen bonding and London forces are key to position and orientation, it is long range electrostatic interactions that control the proximity of the charged surfaces. Rather than the protein surface charge density or surface potential, it is actually the zeta potential at the slipping plane that controls the electrostatic interaction strength.
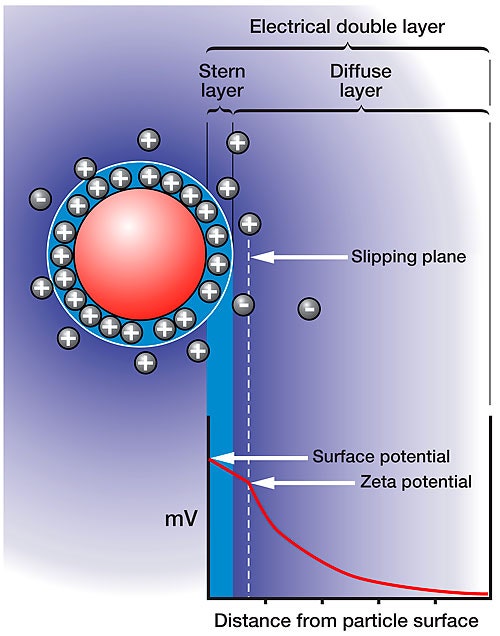
Figure 1 shows a schematic detailing the distribution of ions around a charged particle in solution. The layer closest to the surface of the charged particle is composed of condensed or adsorbed counter ions. The outer boundary of this layer is defined as the Stern layer. A second layer is composed of ions that are less strongly bound, but that have a different balance of ions to the bulk dispersant, called the diffuse layer. There is a notional boundary within this diffuse layer, inside of which the ions move with the particle if the particle moves. This boundary is called the slipping plane, and it is the summation of charge within the slipping plane that defines the strength of the electrostatic interaction, i.e. surface charge + absorbed ions + diffusing ions. The charge or electrostatic potential at the slipping plane is as the definition of the zeta potential.
|
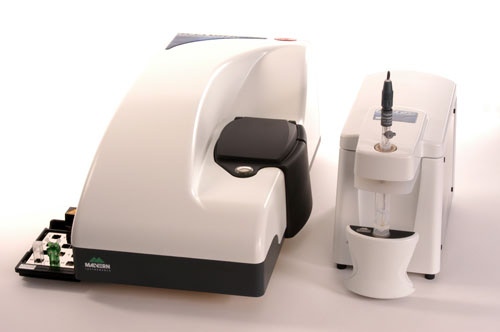
The Zetasizer Nano systems are combined static, dynamic, and electrophoretic light scattering instruments ideally suited for characterization of protein samples. The Zetasizer system measures the size, molecular weight, and zeta potential, and was specifically designed to meet the low sample volume and high sensitivity requirements essential for applications within the field of protein characterization. Automated measurements can be achieved by coupling the Zetasizer system with the MPT-2 autotitrator, which facilitates automated pH, additive, dilution, and conductivity titrations. This application note examines the effects of pH on the size and charge of a typical protein with data collected using a combination of a Zetasizer Nano ZS and MPT-2 autotitrator.
Bovine serum albumin was prepared at 10 mg/ml concentration in 100mM NaCl and adjusted to pH 11. The sample was filtered through a 0.1µm Anotop filter. A 10ml aliquot was transferred to the sample container in the Zetasizer Nano MPT-2 system, which was preloaded with 0.5M HCl, 0.1M HCl, and 0.5M NaOH titrants. The sample was then titrated to pH 2 using increments of 0.5pH units. It is noted here, that while this experiment used 10ml of sample, the MPT-2 titration system can use as little as 3ml sample volume.
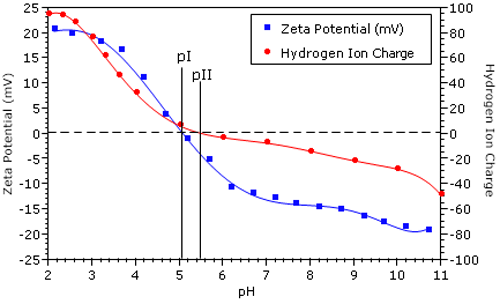
Figure 2 shows the zeta potential vs pH results for a BSA sample in 100 mM NaCl. The isoelectric point (pI = 5.14), defined as the pH of zero electrophoretic mobility, was calculated automatically by the Zetasizer software. Overlaid in Figure 2 is the net protein charge due to hydrogen ion dissociation for BSA in 100 mM NaCl.1 The isoionic point (pII = 5.5) is defined as the pH of net zero H+ charge.
|
The difference between pI and pII is indicative of the specific adsorption of chloride ions. Adsorption of each anionic chloride ion balances one positively charged amino acid residue, leading to a net neutral protein charge at pH less than pII. In the 1950's, Scatchard et al. used membrane dialysis experiments to measure the degree of protein counter-ion binding, and reported that chloride binds very strongly to BSA.2 The results shown in Figure 2 are consistent with those findings.
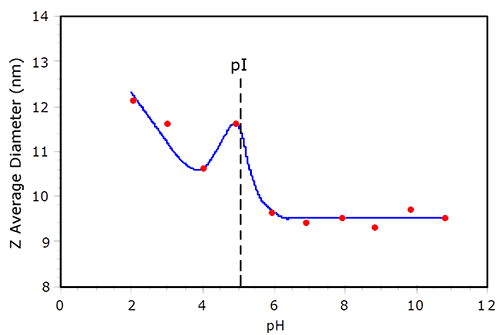
Electrostatic repulsion between like charged molecules is minimized at the isoelectric point (pI), where the net charge at the slipping plane is zero. As the repulsion is minimized, the likelihood of protein aggregation increases. An example of protein aggregation at pI is shown in Figure 3, which shows the pH dependent size of BSA in 100mM NaCl. As the pH is decreased from 11, the hydrodynamic size of BSA remains relatively constant, until the pI is reached. At pH = 5 (~ pI), a local maximum in the size is observed. At pH = 4, the size is smaller than that observed at pI, but then increases as the solution becomes more acidic. In their noted work with BSA, Tanford et al. reported that the protein underwent reversible expansion in the acidic region of the titration curve, with the radius of the equivalent hard sphere at pH 2 being 37% larger than that at pH > pI.3 As seen in Figure 3, the BSA size at pH2 is about 30% greater than the size under basic conditions, consistent with the findings of Tanford et al.
|
The Z average sizes reported in Figure 3 were derived from a Cumulants analysis of the correlation function. Another parameter derived from the Cumulants analysis is the PdI or polydispersity index. While the Cumulants algorithm does not give us the size distribution, the PdI can be used to calculate the width or the polydispersity (Pd) of the distribution. Table 1 shows a comparison of the pH dependent polydispersity for BSA in 100mM NaCl. As seen here, there is very little change in the polydispersity through the course of the titration, except in the region of pI, suggesting that the increase in size under acidic conditions is a consequence of conformational expansion, rather than non-specific aggregation.
| pH | Size (nm) | %Pd |
|---|---|---|
| 10.8 | 9.5 | 45% |
| 9.8 | 9.7 | 48% |
| 8.8 | 9.3 | 46% |
| 7.9 | 9.5 | 46% |
| 6.9 | 9.4 | 45% |
| 5.9 | 9.6 | 47% |
| 4.9 | 11.6 | 58% |
| 4.0 | 10.6 | 48% |
| 3.0 | 11.6 | 47% |
| 2.0 | 12.1 | 47% |
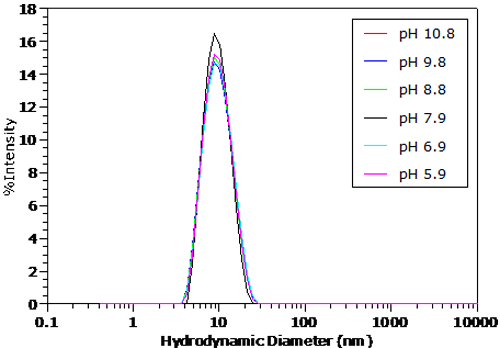
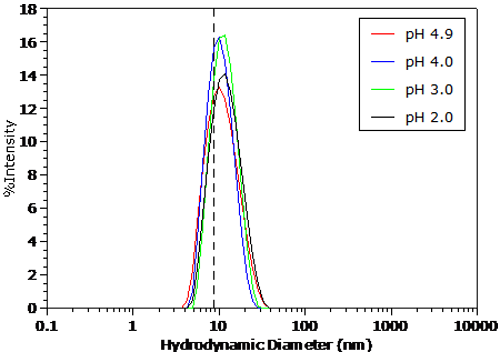
The pH dependent size distributions for BSA in 100mM NaCl are shown in Figures 4 and 5. The distributions were transformed from the measured correlation functions using a non-negative least squares algorithm. The dashed line in Figure 5 indicates the BSA peak position under the conditions in Figure 4. For pH's greater than the pI in Figure 4, no significant change in the measured size distribution is observed. As seen in Figure 5 however, there is a distinct increase in size in the particle size distribution for pH's less than the pI, as suggested by the Z average size results presented earlier.
|
|
|
The optical design of the Zetasizer Nano combined with a versatile autotitrator provides the sensitivity and ease of use to enable the automation of the measurement of biological molecules in a variety of environments. Changes of size and zeta potential can be investigated as a function of pH, ionic concentration, sample concentration and concentration of any soluble additive.