The drug delivery polymer polyglutamic acid was conjugated with different amounts of Doxorubicin and characterized using advanced multi-detector GPC to measure drug content, molecular weight and structure
Work performed in collaboration with the Department of Nanomedicine, Houston Methodist Research Institute, Houston, TX, USA
In pharmaceutical research, correctly targeting the delivery location and release profile of small molecule drugs is a critical challenge. One such strategy being used to overcome this issue is the association of small molecule drugs with larger polymers. This method can be used to control both the drug release profile and the location of drug release, in order to minimize possible side effects.
Doxorubicin (Dox) is a chemotherapy drug that disrupts cell growth and division by inserting itself into the DNA of a cell and interfering with its replication. It is an effective chemotherapy agent, but accurate targeting of the drug's release is difficult. Polyglutamic acid (PG) is a polymer based on the amino acid glutamic acid, which makes it biocompatible. Conjugates of Dox and PG have been shown to effectively kill human triple-negative breast cancer cell lines in vitro[1]. It was shown that the conjugate was effectively taken up by cells and transported to endosomes, where the locally acidic environment cleaves the conjugate, releasing the Dox[1].
In this application note, we show how multi-detector GPC can perform advanced characterization of the PG, Dox and two PG-Dox conjugate samples. This advanced analysis can be used to study drug loading efficiency and the polymer structural changes that occur as the drug is loaded.
PG and PG-Dox conjugates were dispersed in PBS pH 7.4. Dox was dissolved in pure water.
Samples were separated and characterized using OMNISEC, a multi-detector GPC system containing refractive index (RI), UV-Vis (UV), light scattering (LS) and viscometer (IV) detectors. The mobile phase was PBS pH 7.4, containing 30%(v/v) methanol . Malvern A6000M and A3000 columns were used for the separation of aqueous soluble polymers.
The PG and two PG-Dox conjugates could be separated and measured under the specified measurement conditions as shown in Figure 1, with numerical results for PG in Table 1. The PG separated into a single peak, and its weight-averaged molecular weight (Mw) was measured at approximately 13.07 ± 0.08 KDa. The conjugate samples separated into multi-modal peaks with similar retention volumes. The early light scattering peaks (green; 12-14 mL) suggest the presence of some large aggregates. It is also notable that the peaks now contain significant UV absorbance indicating the presence of Dox successfully conjugated to the polymers.
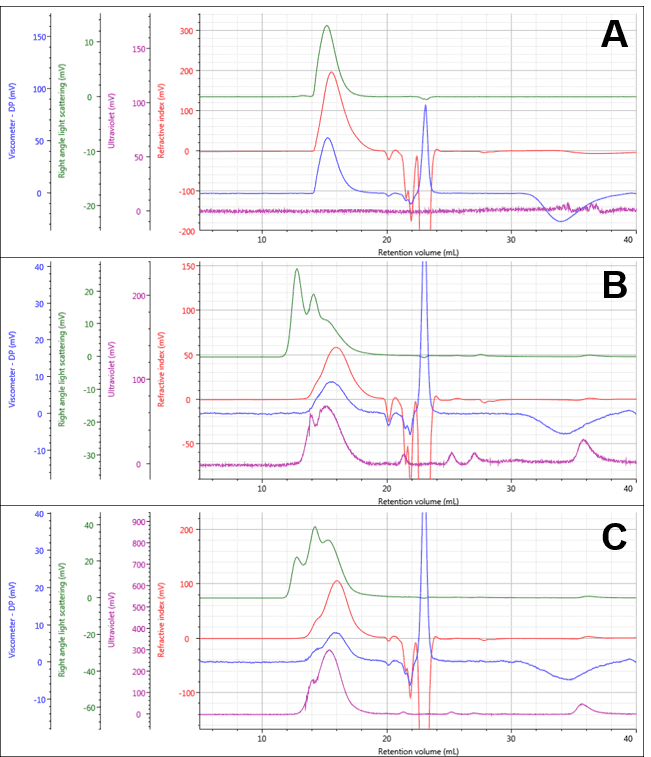
Figure 1: Multi-detector chromatograms of PG (A) PG-Dox Conjugate 1 (B), and PG-Dox Conjugate 2 (C)
| Polyglutamate | |||
|---|---|---|---|
| Mean | SD | % RSD | |
| RV (mL) | 15.58 | 0.009428 | 0.0605 |
| Mn (g/mol) | 7,823 | 96.96 | 1.239 |
| Mw (g/mol) | 13,060 | 72.44 | 0.5545 |
| Mw/Mn | 1.67 | 0.01144 | 0.685 |
| IVw (dL/g) | 0.2696 | 6.61E-04 | 0.2452 |
| Rh(η)w(nm) | 3.679 | 0.003903 | 0.1061 |
| M-H a | 0.717 | 0.00311 | 0.4337 |
| M-H log K (dL/g) | -3.503 | 0.01185 | -0.3382 |
The Dox was also run alone. UV chromatograms of Dox at different mass loadings can be seen in Figure 2A. The free Dox was observed to elute very late from the column, in fact after a full column volume. This clearly indicates significant interaction with the column, delaying the elution of Dox, but method development experiments suggested that despite the interaction, recovery was close to 100%. This is also suggested by the linearity of the concentration response which can be seen in Figure 2B. This calibration curve was used to measure the amount of Dox present in the conjugate samples.
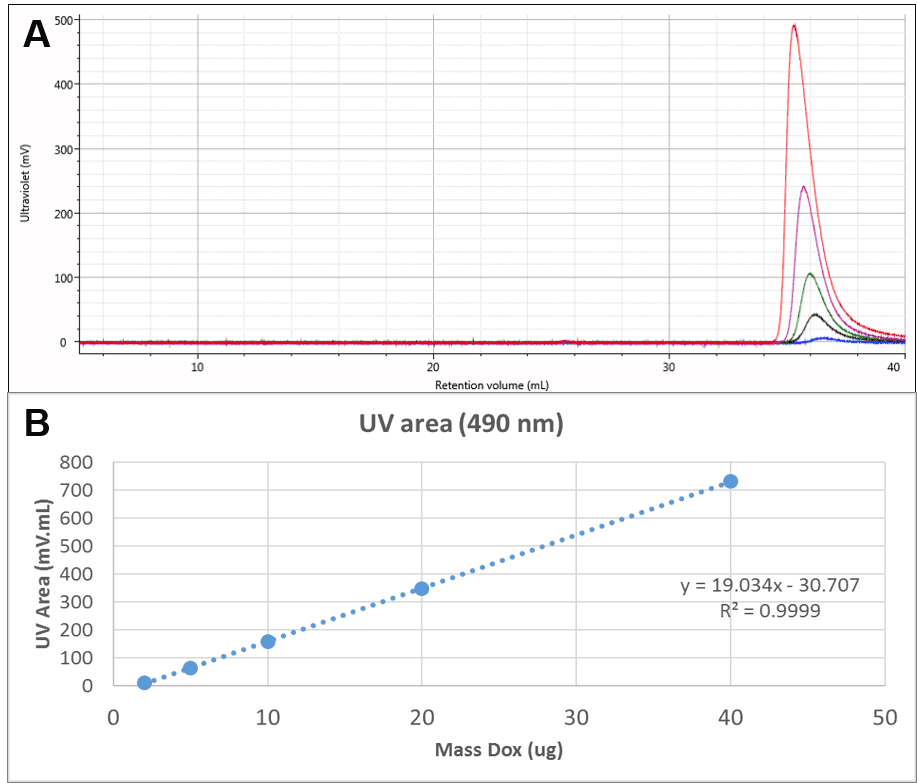
Figure 2: A: Overlaid UV (490 nm) chromatograms of different injection volumes of Dox. B: Concentration calibration curve for Dox
Identifying the peak at 36 mL as free Dox enables this same peak in the PG-Dox conjugate samples to also be identified as free Dox. Comparing the UV spectra for these peaks confirms this conclusion. As can be seen in Figure 3, the conjugate samples can clearly be identified as containing PG-Dox conjugate and free Dox.
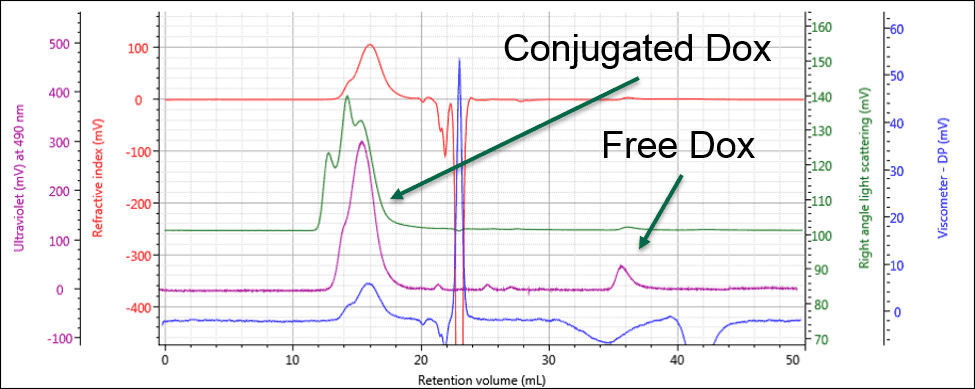
Figure 3: The UV response allows the free and conjugated samples to be identified and quantified
Using the calibration curve from Figure 2B, it is now possible to calculate the amount of Dox present in the conjugate samples.
As shown in Table 2, both PG-Dox conjugates contained a few micrograms of free Dox. Within the injection, PG-Dox 1 contained approximately 11 μg of Dox in the conjugate, while sample 2 contained approximately 39 μg Dox in the conjugate. The total Dox injected and (since injection volume is known) Dox concentration in the sample can then be easily calculated. The approximate total sample concentration can then be calculated from the mass of dissolved material. It is then possible to calculate the approximate loading of Dox in each of the PG-Dox conjugates. It can be approximated from this that sample 2 contained three times as much Dox in the conjugate as did sample 1.
| Conjugate | Dox in main/conjugate peak (µg) | Free Dox (µg) | Total Dox injected (µg) | Dox conc. in sample (mg/mL) | Approx. total conc. (mg/mL) | Approx. fraction Dox (%) |
|---|---|---|---|---|---|---|
| 1 | 11.06 | 3.77 | 14.83 | 0.15 | 2.38 | 6.23 |
| 1 | 11.02 | 4.25 | 15.27 | 0.15 | 2.38 | 6.41 |
| 2 | 38.97 | 4.85 | 43.82 | 0.44 | 2.27 | 19.28 |
| 2 | 39.26 | 5.25 | 44.51 | 0.45 | 2.27 | 19.58 |
Further characterization of the PGA-Dox conjugates is possible but requires a couple of assumptions. During these measurements, it was shown that the Dox fluoresces at the RI and LS wavelength (640 nm). It was therefore not possible to accurately measure the molecular weight of the free Dox. This will also affect the molecular weight measurement of the PG-Dox conjugates, but to a much lesser extent since there is only a small amount of Dox present. Nevertheless, it is possible to make estimates of the PG-Dox conjugate molecular weights using an artificial dn/dc value to offset the effect of the fluorescence. As well as picking an artificial dn/dc, this also assumes that the dn/dc is constant across the peak. Both of these assumptions mean that the results from this analysis will be comparative rather than ‘absolute’. dn/dc values of 0.23 mL/g and 0.34 mL/g were chosen for PG-Dox conjugates 1 and 2, respectively. These values were chosen based on the RI response from the known injection mass. Their artificially high nature (greater than 0.2 mL/g) is a consequence of the fluorescence. These values were used to calculate approximate molecular weight, intrinsic viscosity and structural data for the conjugated polymers, shown in Table 3.
| PG-Dox 1 | PG-Dox 2 | |||||
|---|---|---|---|---|---|---|
| Mean | SD | % RSD | Mean | SD | % RSD | |
| RV (mL) | 15.97 | 0.01414 | 0.08853 | 16.01 | 0.02828 | 0.1767 |
| Mn (g/mol) | 23,900 | 61.94 | 0.2591 | 26,690 | 685.1 | 2.567 |
| Mw (g/mol) | 26,160 | 407.4 | 1.557 | 31,830 | 1,369 | 4.3 |
| Mw/Mn | 1.095 | 0.01988 | 1.816 | 1.193 | 0.02068 | 1.734 |
| IVw (dL/g) | 0.1924 | 0.004384 | 2.278 | 0.1599 | 0.005417 | 3.387 |
| Rh (nm) | 4.267 | 0.05075 | 1.189 | 4.245 | 0.1013 | 2.385 |
| M-H a | 0.6317 | 0.01044 | 1.653 | 0.1859 | 0.009542 | 5.134 |
| M-H log K (dL/g) | -3.499 | 0.04472 | -1.278 | -1.625 | 0.0565 | -3.477 |
The results are known not to be absolute, based on the estimated dn/dc, but the molecular weight and intrinsic viscosity values are in line with expectations and can be seen to broadly represent the changes due to conjugation. This limited accuracy could be addressed in a couple of ways. A light scattering detector with fluorescence filters could be used, such as Malvern’s SEC-MALS 20 FLD. This would exclude the light scattered at wavelengths other than the detector wavelength and would more accurately measure molecular weight. Alternatively, a universal calibration could be performed and the results compared; however, the clear evidence of column interaction suggests this may also have limited accuracy.
As a final step, these results were displayed on a Mark-Houwink (M-H plot). The M-H plot shows intrinsic viscosity as a function of molecular weight and is a visual representation of structural differences between molecules. Polymers that have a higher density in solution appear lower on the M-H plot. It can be used to study structural changes such as branching, or compositional changes such as substitution or conjugation. Figure 4 shows the M-H plots for the 3 samples. Firstly, the two conjugates are significantly lower on the plot than the PG alone. This is expected as the conjugation of the drug molecule will increase the apparent density of the polymers in solution, and the sample with greater Dox loading will move further down. The different slopes between the PG-Dox 1 and 2 suggest that the Dox loading may not be evenly distributed with molecular weight. Further work would be needed to confirm this and to more accurately assess the molecular weight, as this also indicates that the dn/dc will vary with molecular weight. Nevertheless, the plot provides a visual representation of the changes taking place.
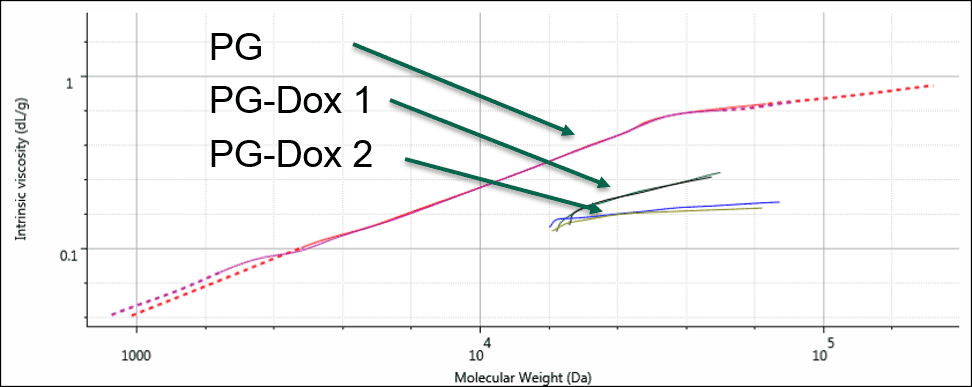
Figure 4: Overlaid Mark-Houwink plots of the PG and PG-Dox conjugate samples
This work shows how multi-detector GPC can be used for advanced characterization of conjugated drug-delivery polymers. Raw polymer, free drug (Dox) and two conjugated products were all characterized under the same measurement conditions. The UV absorbance of the drug enabled the loading levels to be measured and the conjugation levels of the two conjugates to be assessed. With a couple of assumptions, the conjugate molecular weight and structure could also be measured and compared with each other, although in this case, issues with fluorescence prevented the calculation of a completely accurate molecular weight.
There are many challenges associated with the delivery of small molecule drugs. Conjugation with larger polymers is one strategy to improve drug localization and loading, but a high level of characterization is required to achieve reliable and repeatable results. With advanced multi-detector GPC measurements using systems like OMNISEC, scientists can better characterize and understand, and thereby control the amount of drug attached to a polymer delivery vehicle as well as the molecular weight and structure of the product for optimum drug delivery efficiency.