DSC is an established analytical tool used during preformulation and formulation development of biopharmaceuticals. Rank ordering of thermal transition midpoint (TM) by DSC is commonly used to screen formulation buffers, pH and excipients. Typicallly, the higher the TM, the more stable the protein in that formulation, and less likelihood of aggregation. This whitepaper gives examples on the use of DSC in formulation development.
The stability of a candidate therapeutic protein is critical to the success or failure of its development. A protein's stability impacts its production, manufacturing, formulation, long-term storage, delivery to patient, and efficacy. Highly stable proteins tend to encounter fewer issues during manufacturing, are more cost-effective to produce, and have a better chance of remaining functional during formulation and storage. In the “Quality by Design” (QbD) approach to biopharmaceutical development, stability characterization is part of the initial assessment of the “developability” or “drugability” of a candidate molecule,and is constantly revisited during process development and manufacturing. Stability data is also incorporated in the higher order structure (HOS) characterization and “fingerprinting” used for manufacturing support, biopharmaceutical comparability and biosimilarity. Protein HOS characterization is becoming a standard inclusion in regulatory submissions for new biopharmaceutical drugs and biosimilars.
Due to the complex nature of proteins, biophysical characterization tools are important in the analysis of a biopharmaceutical product. There is a variety of biophysical tools which are commonly used to assess protein stability, including (but not limited to) circular dichroism (CD), dynamic and static light scattering (DLS and SLS), size exclusion chromatography–multi-angle light scattering (SEC-MALS), Fourier transform infrared spectroscopy (FTIR), analytical ultrafiltration (AUC), size exclusion chromatography (SEC), differential scanning fluorescence (DSF), intrinsic fluorescence (IF) and differential scanning calorimetry (DSC).
While all of these technologies play an important role in biopharmaceutical development, characterizing thermal stability by DSC is critical. In a 2015 article about biophysical techniques used for monoclonal antibody higher order structure characterization, Gokarn et al. stated: "DSC remains as an unparalleled technique to assess the thermodynamic stability of proteins in a given buffer condition”[1].
This whitepaper focuses on the use of DSC to characterize the thermal stability of protein biopharmaceuticals (primarily antibodies) during preformulation and formulation development, in order to choose the solution conditions in which the protein is preferentially in its native, folded conformation, in order to produce a stable, effective drug product.
DSC is a microcalorimetry technique that is used to characterize the thermal and conformational stability of proteins, nucleic acids, lipids, and other biopolymers [2-7]. DSC measures heat capacity as a function of temperature. The DSC instruments used for protein characterization described in this whitepaper are 'power compensation' instruments, with a fixed-in-place sample cell containing the biopolymer in buffer, and a matched reference cell filled with the same buffer. The heat capacity (Cp) signal from the sample cell is compared with that of the reference cell. As the temperature of the cells is increased, the temperature differences between the reference and sample cells are continuously measured and calibrated to power units. DSC is known a 'forced degradation assay, as the protein is exposed to increasing temperature, the protein begins to unfold, and the Cp of the protein increases correspondingly (Figure 1).
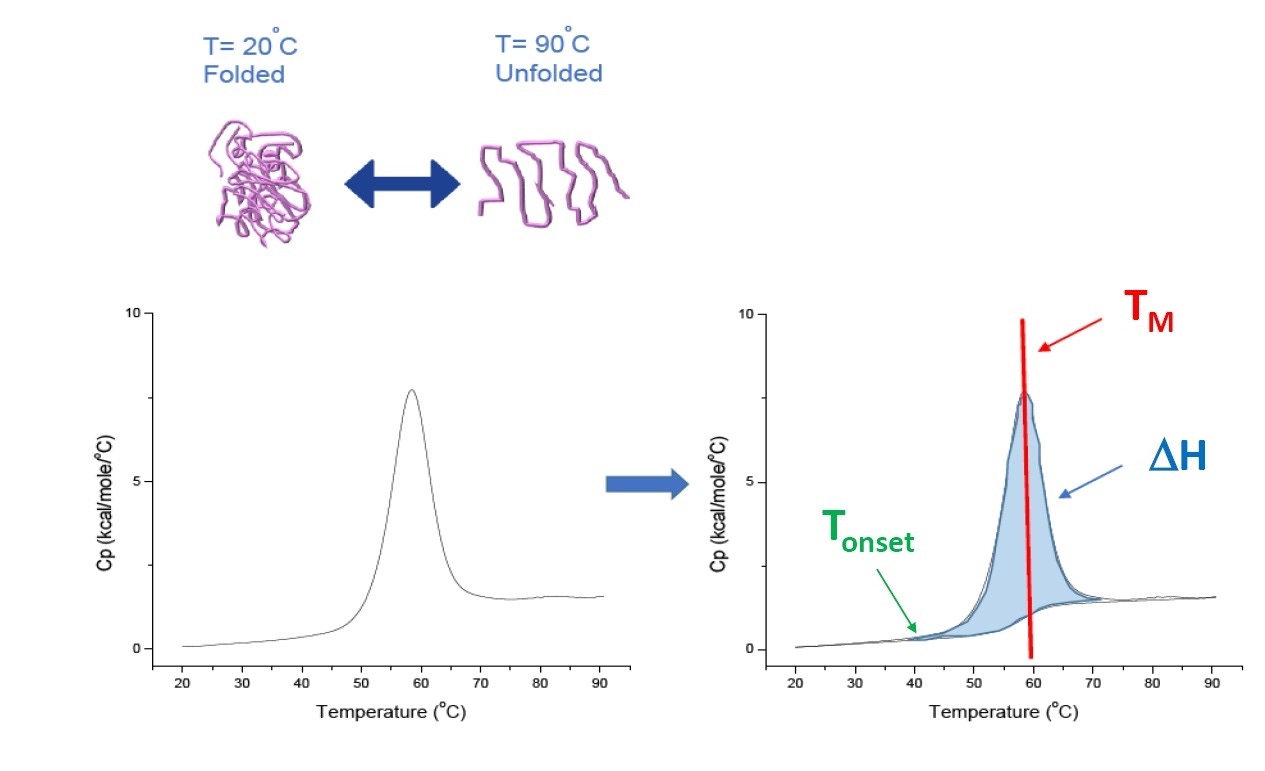
Figure 1: How DSC works: The heat capacity (Cp) changes as a protein thermally denatures. The DSC experiment starts at a temperature at which the protein is primarily folded in its native conformation. With increasing temperature, at some point the protein will begin to unfold/ denature (Tonset) and the Cp increases. At the temperature at which 50% of the protein is in its native conformation and 50% is denatured, the Cp will reach its maximum value - this is the thermal transition midpoint or TM. Above the TM, the protein will be primarily denatured, and at the end of the DSC experiment, all of the protein will be in its unfolded conformation. Experimental parameters for DSC include Tonset, TM, and the unfolding enthalpy (ΔH).
DSC directly measures the heat capacity change, with no need for fluorescence or any other label or probe. For a protein which reversibly denatures, the thermal transition midpoint (TM), also called the melting or denaturation temperature, is the temperature at which the protein is in conformational equilibrium, with 50% in its native (folded) conformation, and 50% in its denatured conformation. The TM is seen as the “peak” of a DSC thermogram.
TM is considered a good indication of thermal stability – the higher the TM, the more thermally stable the protein. Multi-domain proteins (such as antibodies) typically produce more than one peak on a DSC thermogram, so more than one TM can be determined (see Figure 2 for an example).
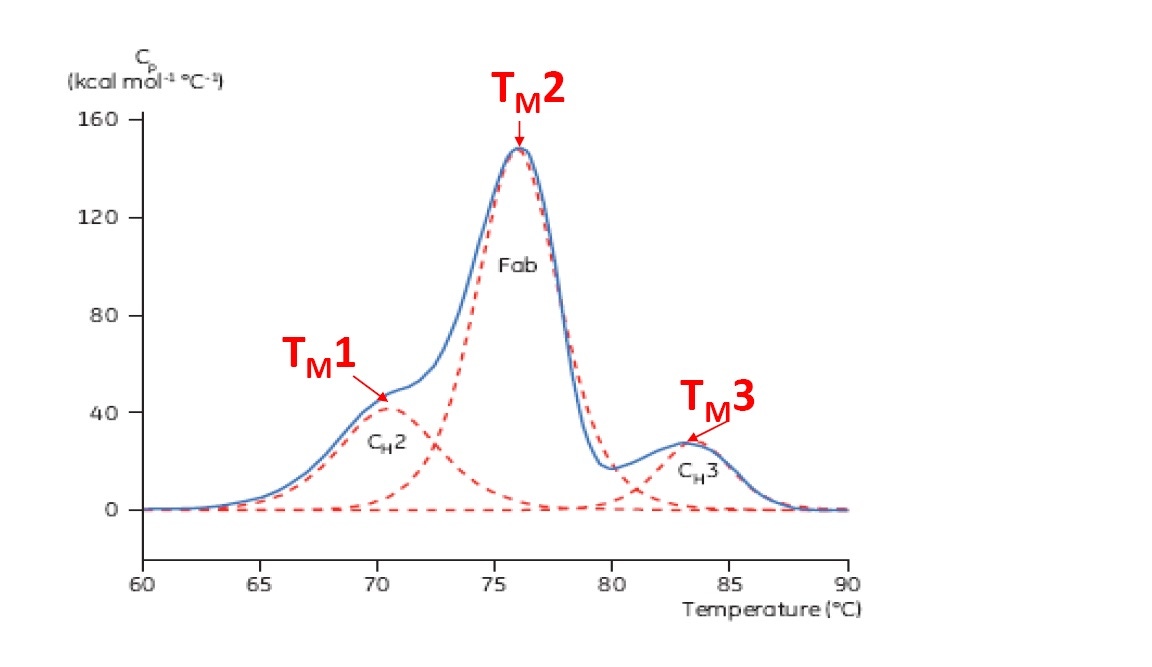
Figure 2: Representative DSC thermogram of a monoclonal antibody, with CH2, Fab, and CH3 domains identified. The dashed red lines are the deconvoluted peaks of each domain transition, with the three TMs indicated.
DSC provides other useful parameters which can be used to characterize and rank protein stability, including the unfolding enthalpy (ΔH), which is measured by the area under the curve. Protein unfolding is an endothermic process, since energy input is needed to break the secondary noncovalent bonds that keep the protein folded correctly. DSC also determines the Tonset (start of unfolding), ΔCp (heat capacity change of unfolding), and T1/2 (width at 1/2 peak height, indicative of the shape of the unfolding thermogram). DSC analysis can include the determination of any combination of these parameters.
Most proteins denature irreversibly, and tend to aggregate or precipitate when heated. The TM and other parameters resulting from the DSC analysis of irreversibly-denatured proteins are not true thermodynamic parameters. However, the rank ordering of TMs from DSC analysis of irreversible protein denaturation is a very useful qualitative parameter for stability screening.
Malvern Instruments offers the MicroCal VP-Capillary DSC system[8,9], which is an automated Differential Scanning Calorimeter designed for TM screening and thermodynamic characterization of proteins and biopolymers in solution.
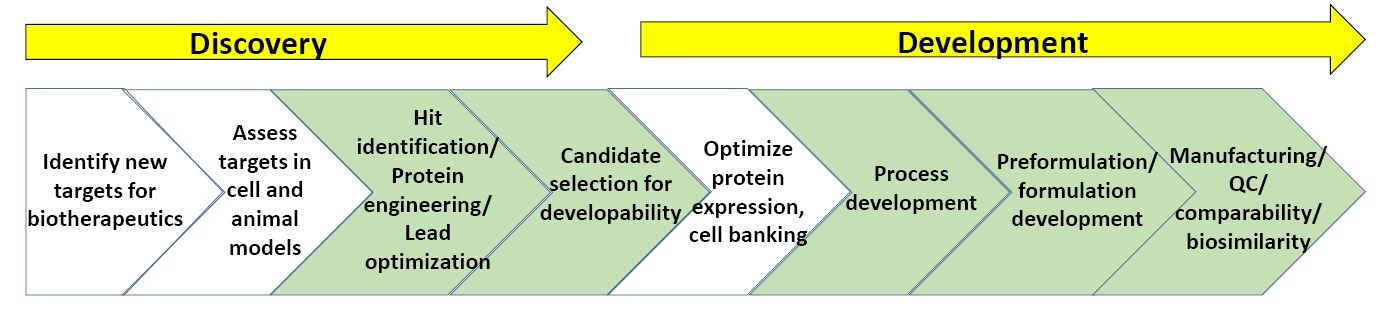
Figure 3. General scheme of processes in biopharmaceutical discovery and development.
Figure 3 shows a generalized scheme of the biopharmaceutical discovery and development pipeline. The sections in green highlight the points at which biophysical characterization, including stability assays, are most commonly utilized (further to this, at the end of this whitepaper is a list of “suggested reading” on biopharmaceutical discovery and development).
The sequence of amino acids which creates a polypeptide chain is referred to as the protein's primary (1°) structure. Beyond this are 3 additional levels of higher order structure (HOS), which it is important to characterize in order to fully understand the stability, functionality, activity and overall nature of the protein. Protein secondary (2°) structure refers to the local folding patterns of a protein’s primary structure, including α-helix, β-sheet, turns and random coils; tertiary (3°) structure refers to the final 3-dimensional structure of a protein, arising from an array of secondary structural elements; finally, the quaternary (4°) structure describes the results of the interaction of two or more identical or different polypeptide chains.
The most common means of administration for biotherapeutics is via the subcutaneous (SC) route. SC-delivered protein drugs must be stable and unaffected at very high protein concentrations (over 100 mg/mL) within their container closures, (e.g. vial or prefilled syringe), often for several years. With that in mind, in order to create a desirable biopharmaceutical, scientists initially seek biomolecules that already demonstrate high stability during candidate selection; however, throughout the development and manufacturing processes, there are many factors that can affect a molecule's stability, meaning that increased stability may need to be introduced via protein engineering.
The purification process involves the removal of the protein from conditions where it is stable, correctly folded, and active, so it is important to carefully adapt buffers, additives, purification methods and storage conditions to keep the protein as stable as possible at this point. When protein molecules are exposed to stressors such as heat, chemicals, pH changes, pressure, mixing, and high concentration, which frequently occurs throughout biopharmaceutical formulation and production, their conformation can favor the denatured (unfolded) form.
Proteins in solution are also susceptible to modifications like deamidation and oxidation, which may lead to denatured, inactive proteins. In the case of protein biopharmaceuticals, denaturation or other modifications could result in the formation of aggregates that lead to products having reduced efficacy or becoming nonfunctional as drugs. Perhaps more significantly, mounting evidence is pointing to the role that protein aggregation may play in creating unwanted and potentially fatal immunogenic responses in patients. Using inherently stable proteins results in more cost-effective production, and successful, effective and safe drug products.
DSC provides a detailed overview of the conformational stability and changes in tertiary and quaternary structure that occur when a protein is thermally denatured, as well as information on how both intrinsic and extrinsic factors can affect protein stability. DSC is considered the best and most quantitative assay for the characterization of thermal stability of biopharmaceutical proteins, and is often used as a predictor of long-term stability[1,10-14]. TMs generated using DSC are a parameter which is frequently used to rank-order stability during candidate selection (developability), formulation screening and process development.Enthalpy (ΔH), Tonset, T1/2, and ΔCpfrom DSC are also used in rank-ordering stability, validation of DSC data, quantitative analysis of protein unfolding, and higher order structure 'fingerprinting'[10-14].
DSC analysis for highly similar proteins in defined solution conditions is reproducible and quantitative (Figure 4). That is, the DSC thermograms will have a similar profile, and parameters (including TM, ΔH and Tonset) will be within an accepted range[12-14]. If the thermograms appear different and the DSC fit parameters change, this suggests that an event such as protein misfolding, degradation, aggregation, differences in buffer, changes in post-translational modification, or other higher order structural change has occurred, impacting the conformational stability of the protein.
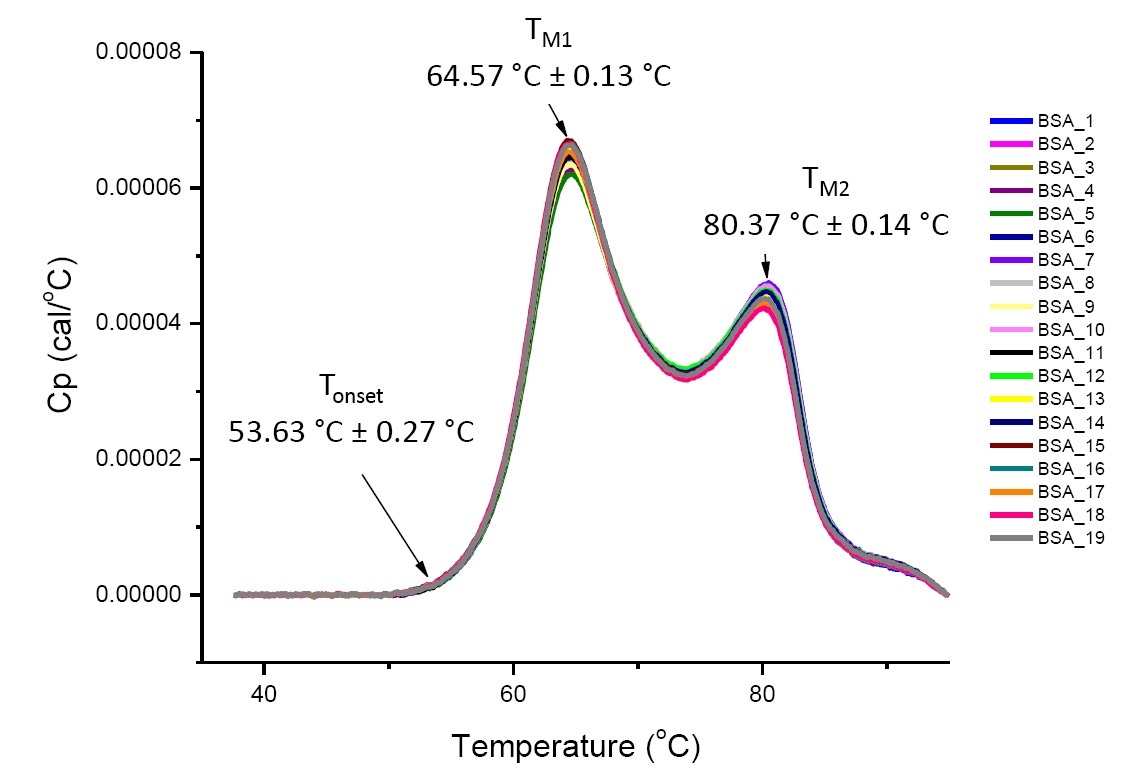
Figure 4: Nineteen DSC thermograms of bovine serum albumin (Sigma A1933, chromatographically purified) in PBS. DSC data shown after scan rate normalization, buffer-buffer subtraction, and integration and baseline subtraction. Mean and standard deviations of Tonset, TM1, and TM2 are shown.
Reproducible and quantitative results make DSC a valuable HOS characterization tool for product evaluation during manufacturing (including batch-to-batch and site-to-site comparability), comparison of protein variants and modified products (including structural changes due to glycosylation, deamidation and oxidation), and biosimilarity. DSC data are also used in regulatory support documents as part of HOS characterization for new drug and biosimilar submissions. In a survey of biopharmaceutical scientists, DSC was ranked as a 'very useful' to 'extremely useful' biophysical analysis technique during candidate selection, formulation development, product characterization, comparability, and biosimilarity[15].
TM values can be determined very simply from DSC thermogram peak(s), without the need for complex data analysis. As we have previously shown (Figure 2), for multi-domain proteins such as antibodies, DSC thermograms show more than one unfolding transition. DSC is able to characterize and quantify the different domains, and can determine the TMs for two, three or more transitions. Other biophysical assays which can determine TM, such as CD, IF and DSF, may only detect the first TM (occurring at the lowest temperature), or the most 'dominant' TM for multi-domain proteins. Extraction of more than one TM from spectroscopic or fluorescent data requires complex data fitting and may not be reproducible.
Compared to other TM screening assays, DSC often requires more protein sample per scan, and can be lower-throughput. If sample is limited, one option would be to perform an initial TM rank ordering with either DSF or IF, and then choose several key samples to validate TM using DSC. It is important that this DSC validation step takes place, and that fluorescence or spectroscopy alone are not relied upon for the measurement of TM in stability assays. Fluorescence-based assays often suffer from artifacts which interfere with measurements, shifting the TM results to a higher (or lower) value. In addition, some proteins and buffer conditions are not compatible with fluorescence, rendering both DSF and IF unsuitable. Finally, fluorescence and spectroscopy are not capable of determining calorimetric enthalpy and other thermodynamic parameters which are supplied by DSC.
DSC is considered by researchers in the biopharmaceutical industry as the 'gold standard' thermal stability assay because the technique:
Measures heat changes associated with protein unfolding
Is a direct measurement of protein unfolding, so no label, probe or tag is needed (as a result, DSC suffers from none of the potential detection artifacts common in fluorescence or other spectroscopic assays)
Is applicable to native proteins in solution
Can be used with practically all buffers and additives in common use for biopharmaceutical purification and formulation. In contrast, many of these buffers and additives are incompatible with fluorescence and spectroscopy
Is easy to set up and operate
Has high precision temperature control, with an operating range up to 130 °C, so can detect most high TM transitions. Most other TM screening assays only heat samples up to 100 °C (or even lower)
Is a 'forced degradation' assay, and does not require storage of protein in heated buffers prior to analysis
Has simple data output and integrated data analysis software
Can be used to resolve separate unfolding transitions and characterize multidomain proteins and protein complexes, as well as simple single-domain proteins
Is information-rich and provides thermodynamic data in addition to conformational stability and TM determination
Can be used as a primary assay for biotherapeutic thermal stability characterization; can also be used with other orthogonal/ complementary biophysical screening tools, and/or validate other data
Is available with high-throughput automation (MicroCal VP-Capillary DSC system) for rapid screening of thermal stability
As a protein-based drug transitions through the pipeline from discovery to development, it is important to develop an appropriate, effective and optimal formulation that maintains its stability, conformation and efficacy during shipping, use, and the full term of its desired shelf life. The formulation must be cost-effective to manufacture. There is a preference for protein formulations that can be self-administered for patient convenience, which requires that the drug product is shipped and stored in a prefilled syringe or similar delivery system that does not require the assistance of a medical professional; in addition, storage of the drug product must be at ambient temperature or in a refrigerator. Some biopharmaceuticals can also be shipped lyophilized (freeze-dried), which requires the drug to be solubilized prior to administration.
Early in the development cycle, sample volume available for testing is often limited, so some companies perform preformulation development, which includes initial small-scale biophysical studies to define the optimal buffer composition and pH to help stabilize the protein. This work helps establish the formulation which will eventually be used to move the protein drug into preclinical and clinical trials. Preformulation development can be conducted at the same time as candidate selection for drug developability assessment, since many of the same biophysical tests are included in both sets of evaluations.
During the preformulation and subsequent formulation development of a protein drug, the protein is exposed to different conditions, including:
Different buffers, pH levels, and salt concentrations
Different formulation additives (excipients). Excipients are inert materials which are present in liquid and lyophilized formulations to help stabilize the protein, or aid in manufacturing or drug delivery, Excipients often used for biopharmaceutical products include surfactants (e.g. polysorbate 80), sugars (e.g. trehalose), polyols (e.g. glycerol), amino acids, preservatives, and anti-oxidants
High protein concentrations, to measure the levels to which a drug candidate may be concentrated (in a range of buffers and additives) before protein aggregation occurs
Extremes of temperature, pressure and humidity
Repeated freeze/thawing cycles and agitation (e.g., in the presence of air/liquid interfaces, transportation stresses)
Contact with a range of different material surfaces, including examples of final container closure (e.g. vial, pre-filled syringe, IV bag)
Different levels of light
Oxidants
Real-time storage tests (to assess the long-term stability potential of the drug product by holding it at intended storage conditions throughout a typical two year shelf life) and accelerated stress studies at elevated temperatures
A 2016 article by Kang et al.[16] evaluated thirty-seven formulations that have been successfully used for commercial monoclonal antibodies. The summary of the evaluation noted:
12 were lyophilized and 25 were liquid formulations (concentrations ranging from 2 mg/mL to 200 mg/mL)
Excipients included: salts, surfactants, polyols, disaccharides, polysaccharides, amino acids, and anti-oxidants
Acetate, citrate, histidine, phosphate, and Tris buffers were commonly used for maintaining optimum pH between 4.7 and 7.4
Most formulations used one of three common surfactants: polysorbate 80 (Tween 80), polysorbate 20 (Tween 20) and poloxamer 188
All lyophilized formulations included sugars (polyols/disaccharides/polysaccharides), while 30% of liquid formulations included sugars
NaCl was commonly used
Glycine and arginine were commonly-used amino acid excipients
A protein in aqueous solution is in equilibrium between its native (folded) and denatured (unfolded) conformations. Hydrophobic interactions and hydrogen bonding are a protein's major stabilizing forces, and these must be overcome in order for the molecule to unfold and denature. Conformational entropy weakens these stabilizing forces, allowing the protein to unfold. In formulation development, the main objective is to find the solution conditions which offer the greatest level of stabilization and therefore enable the highest proportion of native protein. Denatured proteins tend to be more susceptible to irreversible chemical processes such as proteolysis, oxidation, and deamidation, which can in turn lead to inactivation and aggregation.
Biophysical characterization tools are used to monitor protein conformation, predict thermal stability, and measure aggregate formation in response to formulation and storage conditions. Many of the biophysical tools discussed in earlier sections of this whitepaper are included in the preformulation and formulation development workflow. These include DSC, DSF, DLS, SLS, CD, and IF. These assays evaluate protein conformation stability (by TM determination, Tonset, DSC thermogram profile), HOS, particle size and aggregate formation.
Rank ordering of TM by DSC is commonly used as a initial screen of formulation pH, buffers and excipients. Typically, the higher the measured TM, the more stable the protein in formulation (see Figure 5 for DSC thermograms during a pH screen of a protein), and the less likely it is that aggregate formation will occur during storage (see case studies below). DSC is compatible with virtually all buffers and excipients used in biopharmaceutical formulation. Even if a protein's thermal transitions are irreversible in nature, DSC is a convenient and quick method for rank-ordering the effect of buffer conditions and excipients on protein stability throughout formulation development.
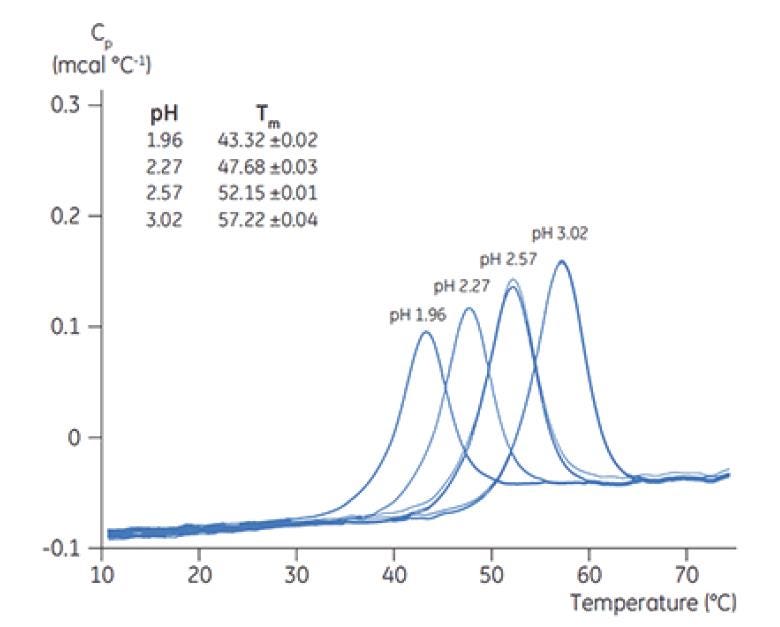
Figure 5. DSC thermograms of chymotrypsinogen at different pH values, with TM indicated for each pH.
MicroCal VP-Capillary DSC systems are used by a significant number of prominent CMOs and CDMOs for the conformational stability characterization and formulation development of biopharmaceuticals. These companies include Patheon[16], Fujufilm Diosynth Biotechnologies[17], and KBI Biopharma[18].
Accelerated stability studies (or 'stress tests') are a common element in formulation screening, and involve storing the candidate molecule in a range of different potential formulations, at elevated temperature(s), and evaluating these over a range of time for conformational stability, size, aggregation, and potential post-translational modifications, using SEC, DSC and DLS.
Once the candidate biopharmaceutical moves further along the development pipeline and there is a sufficient amount of the protein available, the characterization focus turns to the processes involved in final formulation and production, as well as the potential effects of protein concentration and delivery devices. Long-term stability studies are also performed to demonstrate that the protein drug in the chosen formulation has a desirable shelf-life and stability. A good formulation will maintain a biopharmaceutical’s HOS profile for at least two years.
Although there is no absolute guarantee that thermal stability will always correlate with a protein’s physical stability during storage, higher TM does usually indicate a greater energy requirement to unfold the protein. When all other properties of the protein are comparable, the protein formulation(s) which allow better thermal stability are more desirable.
A 1998 publication by Remmele et al.[19] used a MicroCal DSC to characterize the thermal stability of recombinant human interleukin-1 receptor type 1 (IL-1R) in buffers including different excipients. An accelerated stability study (at 30°C and 50°C) using IL-1R stored in solutions from pH 3-9 showed minimal protein breakdown and aggregation at pH 6 (by SDS-PAGE). The DSC thermogram for IL-1R in the 'control' solution (20 mM sodium citrate, pH 6.0) showed the first TMat 48°C, and the second TMat 65.5°C. Twenty-three different excipients were tested, including sugars (mannitol, lactose, sucrose, and glucose), polyols (polyethylene glycol, glycerol and ethanol), surfactants (Tween 80, Pluronic F68), salts (NaCl and CaCl2), and amino acids (lysine, cysteine, alanine, arginine, glycine). The second TMwas not affected by excipients, so the temperature of the first thermal transition was used for rank-ordering of stability. The strategy was to look for excipients that would raise the TMof the low temperature transition, indicating a positive change in native protein stability. Most of the tested excipients resulted in a downward shift in TM(suggesting destabilization of IL-1R), or a small upshift. The additive which created the largest upshift in TM, suggesting improved conformational stability, was 100 mM NaCl, shifting the first TMfrom 48.1°C to 53.1 °C. The authors tried increasing NaCl concentrations from 100 mM to 1500 mM, and continued to see the TMincrease. These results suggested to the authors that the additional thermal stability of IL-1R in the presence of NaCl was due both to the direct interactions of NaCl with the protein, and possibly to changes in water structure. Further excipient analysis was performed in the presence of 100 mM NaCl.
In this same study, the authors looked at the effect that preservatives have on IL-1R stability and aggregation. Preservatives are included in multidose formulations. Using benzyl alcohol, m-cresol and phenol, DSC thermograms showed that all three preservatives destabilized IL-1R , with phenol having the least effect on thermal stability, and benzyl alcohol having the largest destabilizing effect on the TMs. These results were verified by SEC analysis of aggregation formation after 7 and 60 days storage at 37°C – samples including phenol showed the smallest level of aggregation, while those incubated with benzyl alcohol showed the highest level of aggregation.
Remmele and Gombotz[20] showed that measurement of TM by DSC was a good predictor of CD40L aggregation. Performing a pH stability screen with DSC, the highest TMs were measured between pH 6 and pH 7.5, which correlated to the pH range which showed the lowest percentage of aggregation formation as measured by SEC (after storage at 37°C for 7 days).
Remmele[11] summarizes other studies which used data from DSC thermograms to predict and rank-order protein (e.g. chymotrypsinogen and pepsinogen) stability in a variety of formulations. Remmele also discussed how the use of DSC in conjunction with other analytical techniques (SEC-HPLC, CD, AUC, DLS, MS) can provide insight into the conformational stability of the biopharmaceutical in different formulations, information which can then be used to make decisions about which formulation to move forward.
Other formulation screening studies agree that formulations with high TM values (as measured by DSC) tend to demonstrate lower aggregation levels when studied by accelerated stability studies and SEC-HPLC[21-25].
Burton et al.[26] assessed the use of DSC as a rapid screening tool for assessing the stability of two model recombinant antibodies. Changes in TM were monitored as a function of pH and/or excipients, and results were compared with accelerated stability data from samples analyzed by SEC. Data produced by MicroCal DSC correlated with that from SEC (samples with higher TM values measured by DSC also showed the lowest aggregation levels measured by SEC), so DSC was able to determine both optimal solution pH and excipient effects on solution stability. The pH values at which the maximum stability values were predicted by DSC were pH 7.5 for Protein I and pH 6 for Protein II - these values corresponded well with predictions from longer-term solution stability studies for each protein. To quote the authors in the conclusion of the article, “The results of these studies suggest that microcalorimetry can be a valuable tool for rapid screening of protein stability in solution, particularly in the early phase of protein characterization and formulation development when bulk supplies are often quite limited...This can result in significant potential savings in terms of drug substance requirements as well as time and effort spent on sample preparation and complex analysis.” [26]
A Malvern application note highlights how Dr. Katherine Bowers (Fujifilm Diosynth Biotechnologies) utilizes DSC in preformulation development[27]. Antibody X was placed in a range of buffers from pH 3 to pH 8, and DSC analysis (using MicroCal VP-Capillary DSC) was performed immediately (T=0) as well as after storage for 1 week.
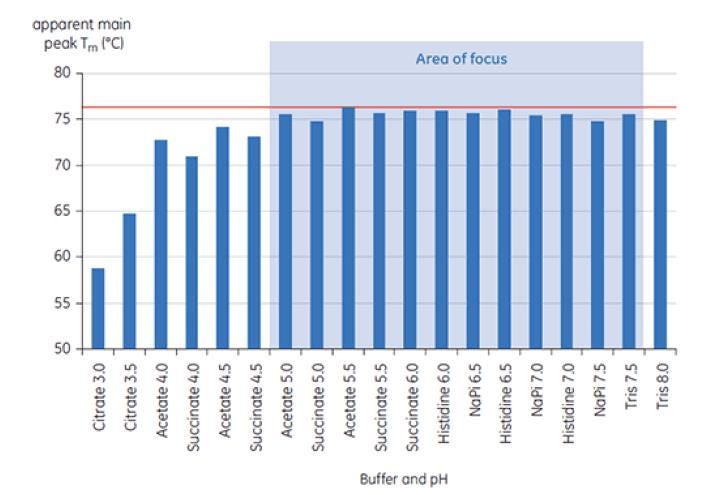
Figure 6: Range of TM values for antibody X in preformulation buffers. Samples were analyzed using the MicroCal VP-Capillary DSC at T= 0.
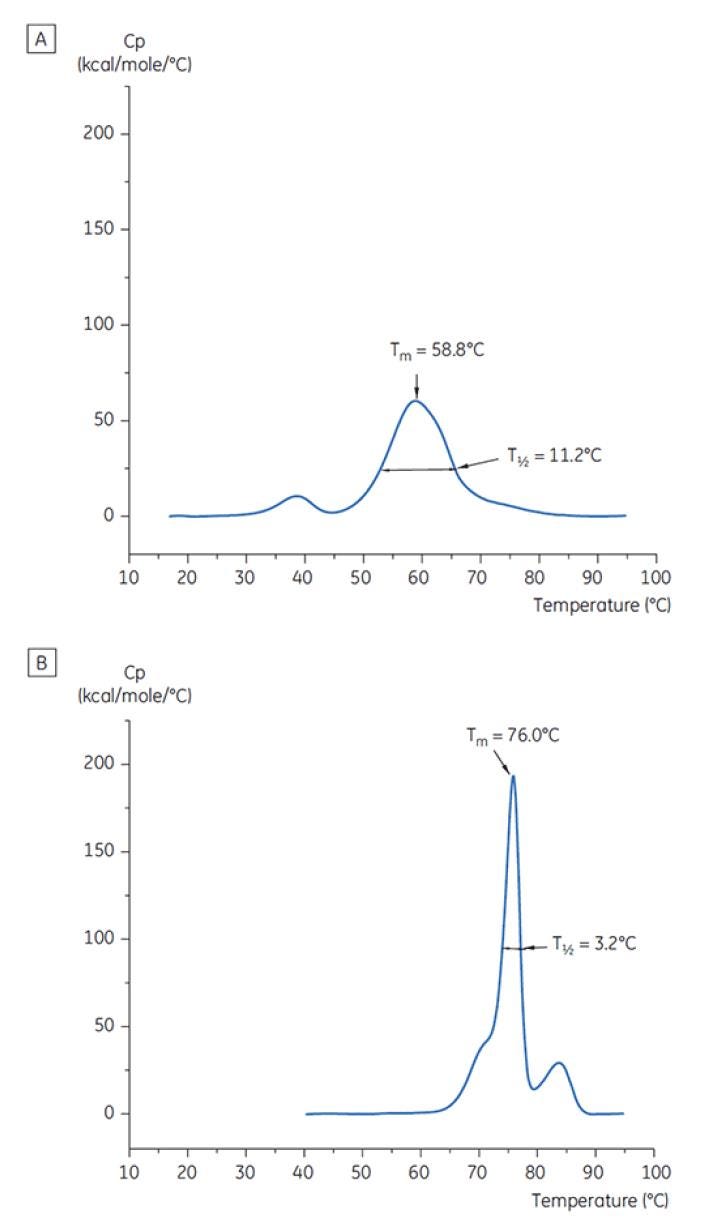
Figure 7: DSC thermograms of antibody X at T= 0. (A) dissolved in citrate buffer, pH 3.0 (B) dissolved in succinate buffer, pH 6.0. TMand T½values are shown for each.
Figure 6 shows the value of the main TM peak at T=0 for antibody X in the initial preformulation screen, and Figure 7 shows the thermogram for antibody X in citrate buffer, pH 3, and succinate buffer, pH 6. From the TM values measured, the most stable buffer conditions (highest TM) were found between pH 5.0 and pH 7.5. At T= 0, other analytical methods (UV, size exclusion chromatography (SEC), light scattering, and SDS-PAGE) showed much less discrimination between the buffer conditions as compared to DSC (data not shown here).
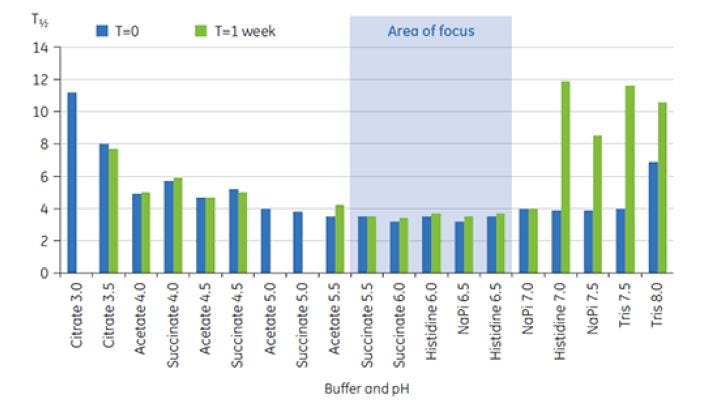
Figure 8: Range of T½values for the main TM transition of antibody X in preformulation buffers at T = 0 and T = 1 week.
T½ values were used to further discriminate between formulation conditions, as indicated in Figure 8. T½ is the peak width at half maximum height for the major transition in the DSC thermogram, and typically reflects the cooperativity of the thermal transition. A lower T½ value can indicate a more compact structure and is therefore preferred for formulations. Here, the lowest T½ values after storage for one week were found for buffers with pH values between 5.5 and 6.5 (Figure 8). These data were used to rank appropriate buffers and pH interval for the subsequent excipient screening, considerably reducing the number of exploratory conditions. This study shows that DSC can be used to quickly optimize pH and buffer conditions for preformulation development.
A biopharmaceutical product is also characterized in different buffers and excipients to understand the potential protein degradation pathway(s). This information aids in developing the final formulation. Typically, biopharmaceutical drugs do not degrade when stored in a refrigerator during formulation development, so a temperature-stress accelerated stability study is usually performed, as discussed above in several examples. A study by Zheng et al.[28] examined the degradation of a IgG1 therapeutic monoclonal antibody mAb-A. This group first evaluated mAb-A in different buffers (sodium phosphate, sodium citrate, and sodium acetate), different NaCl concentrations (20 mM and 100 mM), and different pH values (pH 4.5, 5.5, 6.5, and 7.5), using a MicroCal VP-Capillary DSC. The most promising formulation candidates from each buffer group, based on the highest TM and Tonset values, were selected for further evaluation: sodium phosphate pH 6.5 plus 20 mM NaCl, sodium citrate pH 6.5 plus 20 mM NaCl, and sodium acetate pH 6.5 plus 20 mM NaCl. mAb-A in these three buffers showed similar TMs and Tonsets, as well as nearly identical DSC thermogram profiles. As a comparison, this study also examined a formulation in sodium acetate pH 4.5 plus 20 mM NaCl, which has a lower thermal transition and TM.
The authors stored mAb-A in the four formulations at 40°C, and the proteins were further characterized by SEC (to separate degradation products from native protein), DLS (to determine the modified second virial coefficient in order to characterize protein-protein interactions), LC-MS (to sequence the degradation products), hydrophobic interaction chromatography (to separate degradation products from native protein), and SDS-PAGE (to determine the molecular weights of degradation products). It was found that storage in pH 4.5 buffer caused more fragmentation compared with storage in the same buffer at pH 6.5. In addition, storage at pH 4.5 induced the unfolding of the CH2 domain, increasing the surface accessibility of mAb-A, which may have facilitated fragmentation. Although DSC showed that the conformational structure of mAb-A in phosphate pH 6.5, citrate pH 6.5, and acetate pH 6.5 buffers were similar, DLS showed that the potency of the protein-protein interaction was different in each of the buffers. Interpreted together, the results from this study suggested that protein-protein interactions played the major role in controlling aggregate formation in mAb-A. The authors also stated that the knowledge obtained from this study was useful for mAb-A formulation development, but should not be extrapolated to other mAbs.
Proteolysis of biopharmaceuticals during purification and storage can also be an issue. Host cell proteases may affect the quality of recombinant proteins and can also cause significant product loss. Unfolded or partially-folded proteins are more prone to proteolysis than folded proteins, and folded proteins can be stabilized against proteolysis by the addition of osmolytes.
In a study by Mueller, et al.[29], the mechanism of protection of an immunoglobulin M (IgM) from proteolysis was investigated. IgMs are emerging as therapeutic candidates, but they have the reputation of being inherently unstable. In this study, the proteases pepsin, papain, and α-chymotrypsin were added to a purified IgM, mAb 85, and the degradation was analyzed in the absence and presence of excipients. An increase in conformational stability of mAb 85 as a result of adding the excipients glycine and sorbitol was confirmed using MicroCal VP-Capillary DSC. mAb 85 unfolded in 2–3 thermal transitions dependent on the pH of the buffer, these transitions corresponding to different domains or structural areas. The TMs were significantly increased by adding 20% sorbitol or 1 M glycine (increase of 4–5°C) or both (increase of 7–11°C) at pH 5.5 and pH 7.4. This raises the question of how the protection by sorbitol and glycine is occurring. One possibility is that the proteases are inhibited by the presence of these excipients, perhaps by compaction of their standard conformations. Another possibility is compaction of the IgM, making it less accessible for proteolytic cleavage.
To elucidate the mechanism, the proteolytic cleavage of low molecular weight substrates was tested. These substrates are not stabilized by compaction of their conformation via the preferential exclusion mechanism. The activity of papain was slightly increased, while the activity of chymotrypsin was virtually unchanged, suggesting that the excipients might stabilize papain without reducing its activity. These results indicate that the protective effect of the excipients is mediated solely through the conformational stabilization of mAb 85, which may include increased compaction and a possible reduction in the accessibility of the cleavage sites for proteases. The addition of excipients such as sorbitol and glycine to buffers during purification and formulation may therefore help reduce or even eliminate the requirement for protease inhibitors in biopharmaceutical formulations.
An example of how DSC and other biophysical methods can be used in formulation development specific to a particular delivery system is shown in an article by Morar-Mitrica et al. [30]. Otelixizumab is a humanized mAb (IgG1) directed against human CD3. Since the clinical doses are low (0.1 mg to 0.5 mg per dose), the drug product was developed at a concentration of 0.2 mg/mL. The dose administration required dilution of the mAb in an IV bag containing normal saline, followed by delivery of the contents of the entire bag via infusion pump. This led to protein concentrations in the IV bag as low as 0.002 mg/mL. Due to the low protein concentration, there was a high risk of significant protein loss by interaction with the IV bag and/or the infusion pump system. Through conventional biophysical formulation development and in-use stability studies, a drug product was developed which had reduced adsorptive losses and reduced oxidative degradation, and the liquid formulation was stable under refrigerated conditions.
For standard formulation screening, TM values for otelixizumab in the presence and absence of 0.1% polysorbate 80 (PS80) were determined from DSC thermograms. Thermograms of protein in the presence and absence of PS80 were similar, with a consistent unfolding profile, suggesting that there were no HOS changes as a function of surfactant. At least two species/domains, corresponding to 2 unfolding transitions, were clearly identified by DSC. The DSC results for the mAb in the absence and presence of PS80 showed no significant differences in either TM1 and TM2. In addition, PS80 had no effect on the thermal stability of the mAb, as indicated by a comparison of other relevant DSC-derived thermodynamic parameters such as T1/2 and total enthalpy of unfolding.
Peroxides can be present in significant quantities in raw materials such as polysorbates, causing potential oxidative damage to biopharmaceutical products, and resulting in consequences such as reduced efficacy, and possibly in unwanted immunogenic response. To assess the risk of oxidation as a significant degradation pathway, otelixizumab in histidine buffer alone (containing no PS80) was oxidized by treatment with hydrogen peroxide, and the oxidation-induced structural changes were evaluated by DSC and compared to a non-oxidized control. TM analysis showed an oxidation-induced destabilization of the first unfolding transition, typically assigned to the CH2 domain of the mAb. There was a significant decrease in TM1 (the low-temperature transition) concomitant with peak broadening. The second transition (TM2) was not affected by oxidation. Mass spectrometry (MS) analysis confirmed that one exposed methionine residue in the CH2 domain was 90% oxidized by peroxide. These results demonstrated that the major oxidation site was the methionine residue in the CH2 domain and that the oxidation event correlated with the thermodynamic destabilization of this domain. The authors concluded that the administration of low concentration therapeutic mAbs has challenges, and it is critical that formulation development and in-use stability studies are performed early in development.
Results presented in this whitepaper clearly demonstrate the importance and effectiveness of incorporating DSC as a biophysical stability assay during preformulation and formulation development. Using DSC results, along with other stability assays, biopharmaceutical companies can make informed decisions on the most stable formulations, meaning that the protein is less likely to exhibit long-term stability and aggregation issues in its final formulation and as a drug product. This translates into more cost-effective drug production, and an increased likelihood that the final drug formulation will remain active, stable, safe, and in the correctly-folded conformation.
Analytical Techniques for Biopharmaceutical Development, R. Rodriguez-Diaz, T. Wehr, S. Tuck (eds.), Taylor & Francis, New York USA (2005).
Biophysical Characterization of Proteins in Developing Biopharmaceuticals, D.J.Houde, S.A. Berkowitz (eds.), Elsevier, Amsterdam, Netherlands (2015).
Biophysical Methods for Biotherapeutics: Discovery and Development Applications, T.K. Das (ed.) John Wiley & Sons, Hoboken NJ USA (2014).
Biophysics for Therapeutic Protein Development, L.O. Nahri (ed.), Springer NewYork, USA (2013).
State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal AntibodyCharacterization Volume 1. Monoclonal Antibody Therapeutics: Structure, Function, and Regulatory Space, J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1176 (2014). doi: 10.1021/bk-2014-1176.
State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 2. Biopharmaceutical Characterization: The NIST mAb Case Study, J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1201 (2015). doi: 10.1021/bk-2015-1201.
State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 3. Defining the Next Generation of Analytical and Biophysical Techniques, J.E. Schiel, D.D. Davis, O.V. Borisov (eds.), ACS Symposium Series Vol 1202 (2015) DOI: 10.1021/bk-2015-1202.