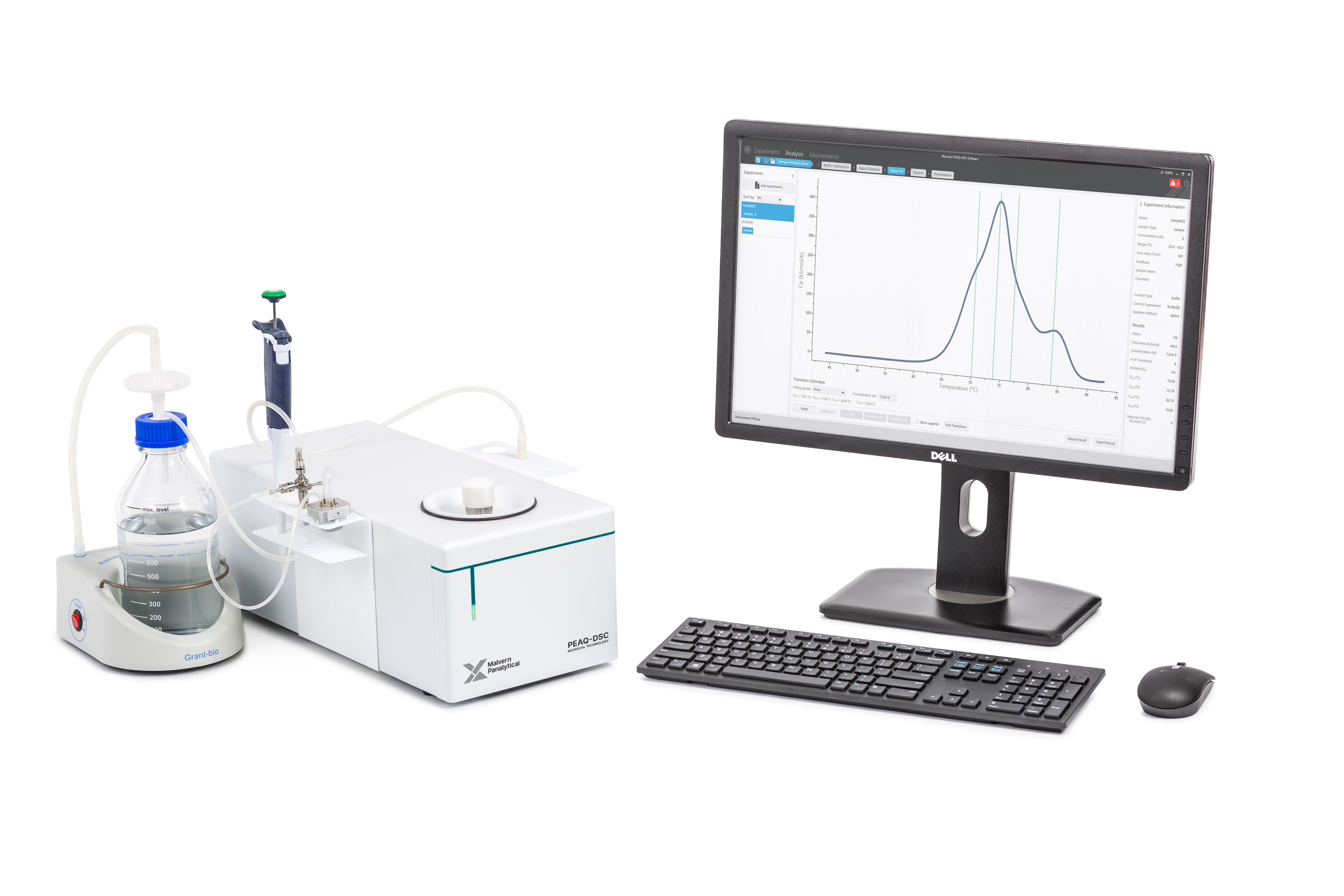


The PEAQ-DSC system, a MicroCal technology, provides highly sensitive, easy-to-use microcalorimetry that helps reduce the time and cost associated with stability testing and comparability analysis. The PEAQ-DSC is a manual instrument with a cleaning device, which can be upgraded to an automated version upon request.
Select a model:
Looking for more information?
To request a quote, more information or download a brochure select an option below.
The high-sensitivity MicroCal PEAQ-DSC system features automated data analysis to support the generation of high-integrity thermal stability data and deliver compliance with regulatory requirements, whilst allowing easy integration into existing data handling and transfer systems.
Unattended operation of the instrument after sample loading frees operator time. This complete system requires no additional accessories, reagents or consumables.
The PEAQ-DSC Automated, with an autosampler, is also available.
Gold standard stability-indicating technique requiring minimal assay development
Direct and label-free quantification of biomolecular native-state stability in solution
Measurement of very tight binding constants, up to 1020 M-1
Manual system with pipette and cleaning device
Powerful MicroCal PEAQ-DSC software minimizes typical data analysis time
Data generated by the MicroCal PEAQ-DSC provides critical guidance for biopharmaceutical development in protein engineering, (pre)formulation development, process development, manufacturing change control and biosimilarity and biocomparability studies.
The integrated software streamlines workflows, facilitates non-subjective data analysis, performance qualification and compliance with 21 CFR Part 11 and Annex 11 regulations, all delivering high integrity data and driving productivity in biopharmaceutical research.
Powerful MicroCal PEAQ-DSC software minimizes typical data analysis time and includes:
Differential Scanning Calorimetry (DSC) is a powerful analytical tool for characterizing the thermal stability of proteins and other biomolecules. The technique measures the enthalpy (ΔH) and temperature (Tm) of thermally-induced structural transitions of molecules in solution. This information provides valuable insights into factors that stabilize or destabilize proteins, nucleic acids, micellar complexes and other macromolecular systems.
The data are used to predict shelf-life of biomolecular products including biopharmaceuticals, to enable batch-to-batch and biosimilar versus innovator molecule comparisons, to develop purification strategies, to characterize and evaluate protein constructs, and to rank the affinities of ligands to their protein targets in small molecule drug discovery programs.
Data analysis and measured/derived parameters are described in greater detail on the Differential Scanning Calorimetry technology page.
Functional structures formed by proteins and other macromolecules often undergo temperature-induced conformational changes, such as unfolding. These changes result in the absorption of heat as a result of the redistribution of non-covalent bonds within the molecule. Differential Scanning Calorimeters measure this heat uptake very precisely.
The thermal core of the PEAQ-DSC system contains a sample cell (containing the sample of interest) and a reference cell (containing a matched buffer solution), both of which are housed within an insulating jacket. These two cells are always maintained at the same temperature, and during a measurement, they are heated at a constant scan rate.
As the molecule within the sample cell unfolds, heat is absorbed, creating a temperature difference (ΔT) between the sample cell and the reference cell. This results in a thermal gradient across the Peltier units, creating a proportional voltage, which is in turn converted to power to form a feedback loop to the Peltier units, in order to return ΔT to zero.
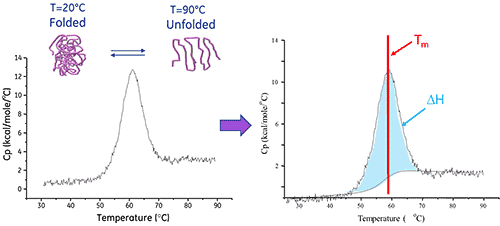
Since protein unfolding is an endothermic event, it is observed as a positive displacement in the thermogram. The midpoint of this protein ‘melting’ transition is the Tm, and the area under the curve is the enthalpy (ΔH) of the unfolding process.
MicroCal PEAQ-DSC instruments provide value to a wide range of applications, some of which are outlined below.
MicroCal PEAQ-DSC is a fundamental tool for characterizing proteins. Key questions such as below can all be answered in simple DSC experiments.
Liposomes are used as model membrane systems and as potential drug delivery systems. ‘Structure Activity Relationships’ (SAR) of membrane components and drug uptake into a liposomes can be investigated monitoring shifts in the phase transition temperatures and enthalpies measured by DSC. Indeed, any material that forms micelles or other macromolecular structures can be studied using the technique.
Application note: Using DSC to characterize thermotropic phase transitions in lipid bilayer membranes: The basics of liposome sample preparation and DSC studies
| Technology | Differential Scanning Calorimetry |
|---|---|
| Measurement type | Temperature midpoint (TM)
Enthalpy (ΔH)
Heat capacity change (ΔCp) |
| Cell | Capillary |
|---|---|
| Cell material | Tantalum |
| Cell volume | 130 μL |
| Sample volume | 250 µL (manual filling) |
|---|---|
| Typical sample concentration | 0.01 mg/mL – 10 mg/mL 1 |
| Sample throughput | ≤ 6 samples analyzed/8h |
| Noise | 0.05 μCal/°C 2 |
|---|---|
| Baseline repeatability | 1 μCal/°C 2 |
| Response time | 5 s2 |
| Repeatability | <0.2 µCal/°C 3 |
| Reproducibility | <0.08°C St. Dev. TM and <2% RSD on ΔH 4 |
| System reproducibility | <0.1°C St. Dev. TM and <5% RSD on ΔH 4 |
| Multiple feedback modes | Yes (passive, high gain and low gain) |
| Temperature range | 2 - 130 °C 2,5 |
| Maximum scan rate | 240 °C/h |
| Reverse scanning | Yes |
| Pressure perturbation calorimetry (PPC) | N/A - manual cleaning device with guided workflows |
| Cleaning solvents | Water and detergent (Contrad 90) used as standard |
| 21 CFR part 11 | Yes, with PEAQ-Compliance software option |
|---|---|
| Network ready | Yes, with email alert capability |
| Operating temperature (°C) | +10 °C to +28 °C |
|---|---|
| Storage temperature | -20 °C to +50 °C |
| Humidity | 10% to 70%, non-condensing (10% to 90% for storage) |
| Ingress Protection (IP) rating | IP21 |
| Power | 100-240 V A/C, 50/60 Hz, 70 W (cell), PC as supplied |
| Certification | CE (EN61010-1), EMC (EN61326-2-1, EN61326-1, FCC, ICES, VCCI), ISO9001:2008 |
| Dimensions (W, D, H) | 20 cm × 44 cm × 19 cm |
|---|---|
| Weight | 8.2 kg |
1 Sample dependent
2 Typical results for ribonuclease (RNase) in 50 mM KAc buffer at pH 5.5, at 60 °C/h with high feedback 3 Rescans of a stable buffer 4 Using ribonuclease (RNase) 5 Range may be extended down to -10 °C upon request |
Version number: MAN0587-02-EN-00
Version number: MAN0588-02-EN-00
MicroCal PEAQ-DSC Software Update v1.65 - Maintenance release
MicroCal PEAQ-DSC Software Update v1.64 - Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Initial release of MicroCal PEAQ-DSC software
|
|
Unmatched uptime and performance
Malvern Panalytical instruments are built to last and good maintenance and care ensure continued peak performance for many years to come. Service Agreements build this into your schedule and budget, based on preventive maintenance and remote support . Different levels include repair visits, response times and parts coverage as needed. OQ services, training and application consultancy can be added as well.
|
|
|
|
Advanced analysis and actionable insights
|
|
|
|
Easy start and streamlined routine operations
|
|
|
|
Want to speak to a service expert or find the services that are just right for you?
Reach out to our experts and evaluate your services needs and business risks by answering a few questions around the usage of your Malvern Panalytical instrument.
|
|
|
*Please note that the exact details of our services and their availability are determined by a variety of customer-specific factors, such as product type, product configuration, or location. |
||
The high-sensitivity MicroCal PEAQ-DSC system features automated data analysis to support the generation of high-integrity thermal stability data and deliver compliance with regulatory requirements, whilst allowing easy integration into existing data handling and transfer systems.
Unattended operation of the instrument after sample loading frees operator time. This complete system requires no additional accessories, reagents or consumables.
The PEAQ-DSC Automated, with an autosampler, is also available.

Gold standard stability-indicating technique requiring minimal assay development
Direct and label-free quantification of biomolecular native-state stability in solution
Measurement of very tight binding constants, up to 1020 M-1
Manual system with pipette and cleaning device
Powerful MicroCal PEAQ-DSC software minimizes typical data analysis time

Data generated by the MicroCal PEAQ-DSC provides critical guidance for biopharmaceutical development in protein engineering, (pre)formulation development, process development, manufacturing change control and biosimilarity and biocomparability studies.
The integrated software streamlines workflows, facilitates non-subjective data analysis, performance qualification and compliance with 21 CFR Part 11 and Annex 11 regulations, all delivering high integrity data and driving productivity in biopharmaceutical research.
Powerful MicroCal PEAQ-DSC software minimizes typical data analysis time and includes:

Differential Scanning Calorimetry (DSC) is a powerful analytical tool for characterizing the thermal stability of proteins and other biomolecules. The technique measures the enthalpy (ΔH) and temperature (Tm) of thermally-induced structural transitions of molecules in solution. This information provides valuable insights into factors that stabilize or destabilize proteins, nucleic acids, micellar complexes and other macromolecular systems.
The data are used to predict shelf-life of biomolecular products including biopharmaceuticals, to enable batch-to-batch and biosimilar versus innovator molecule comparisons, to develop purification strategies, to characterize and evaluate protein constructs, and to rank the affinities of ligands to their protein targets in small molecule drug discovery programs.
Data analysis and measured/derived parameters are described in greater detail on the Differential Scanning Calorimetry technology page.
Functional structures formed by proteins and other macromolecules often undergo temperature-induced conformational changes, such as unfolding. These changes result in the absorption of heat as a result of the redistribution of non-covalent bonds within the molecule. Differential Scanning Calorimeters measure this heat uptake very precisely.
The thermal core of the PEAQ-DSC system contains a sample cell (containing the sample of interest) and a reference cell (containing a matched buffer solution), both of which are housed within an insulating jacket. These two cells are always maintained at the same temperature, and during a measurement, they are heated at a constant scan rate.
As the molecule within the sample cell unfolds, heat is absorbed, creating a temperature difference (ΔT) between the sample cell and the reference cell. This results in a thermal gradient across the Peltier units, creating a proportional voltage, which is in turn converted to power to form a feedback loop to the Peltier units, in order to return ΔT to zero.
Since protein unfolding is an endothermic event, it is observed as a positive displacement in the thermogram. The midpoint of this protein ‘melting’ transition is the Tm, and the area under the curve is the enthalpy (ΔH) of the unfolding process.

MicroCal PEAQ-DSC instruments provide value to a wide range of applications, some of which are outlined below.
MicroCal PEAQ-DSC is a fundamental tool for characterizing proteins. Key questions such as below can all be answered in simple DSC experiments.
Liposomes are used as model membrane systems and as potential drug delivery systems. ‘Structure Activity Relationships’ (SAR) of membrane components and drug uptake into a liposomes can be investigated monitoring shifts in the phase transition temperatures and enthalpies measured by DSC. Indeed, any material that forms micelles or other macromolecular structures can be studied using the technique.
Application note: Using DSC to characterize thermotropic phase transitions in lipid bilayer membranes: The basics of liposome sample preparation and DSC studies
| Technology | Differential Scanning Calorimetry |
|---|---|
| Measurement type | Temperature midpoint (TM)
Enthalpy (ΔH)
Heat capacity change (ΔCp) |
| Cell | Capillary |
|---|---|
| Cell material | Tantalum |
| Cell volume | 130 μL |
| Sample volume | 250 µL (manual filling) |
|---|---|
| Typical sample concentration | 0.01 mg/mL – 10 mg/mL 1 |
| Sample throughput | ≤ 6 samples analyzed/8h |
| Noise | 0.05 μCal/°C 2 |
|---|---|
| Baseline repeatability | 1 μCal/°C 2 |
| Response time | 5 s2 |
| Repeatability | <0.2 µCal/°C 3 |
| Reproducibility | <0.08°C St. Dev. TM and <2% RSD on ΔH 4 |
| System reproducibility | <0.1°C St. Dev. TM and <5% RSD on ΔH 4 |
| Multiple feedback modes | Yes (passive, high gain and low gain) |
| Temperature range | 2 - 130 °C 2,5 |
| Maximum scan rate | 240 °C/h |
| Reverse scanning | Yes |
| Pressure perturbation calorimetry (PPC) | N/A - manual cleaning device with guided workflows |
| Cleaning solvents | Water and detergent (Contrad 90) used as standard |
| 21 CFR part 11 | Yes, with PEAQ-Compliance software option |
|---|---|
| Network ready | Yes, with email alert capability |
| Operating temperature (°C) | +10 °C to +28 °C |
|---|---|
| Storage temperature | -20 °C to +50 °C |
| Humidity | 10% to 70%, non-condensing (10% to 90% for storage) |
| Ingress Protection (IP) rating | IP21 |
| Power | 100-240 V A/C, 50/60 Hz, 70 W (cell), PC as supplied |
| Certification | CE (EN61010-1), EMC (EN61326-2-1, EN61326-1, FCC, ICES, VCCI), ISO9001:2008 |
| Dimensions (W, D, H) | 20 cm × 44 cm × 19 cm |
|---|---|
| Weight | 8.2 kg |
1 Sample dependent
2 Typical results for ribonuclease (RNase) in 50 mM KAc buffer at pH 5.5, at 60 °C/h with high feedback 3 Rescans of a stable buffer 4 Using ribonuclease (RNase) 5 Range may be extended down to -10 °C upon request |
Version number: MAN0587-02-EN-00
Version number: MAN0588-02-EN-00
MicroCal PEAQ-DSC Software Update v1.65 - Maintenance release
MicroCal PEAQ-DSC Software Update v1.64 - Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Maintenance release
Initial release of MicroCal PEAQ-DSC software
|
|
Unmatched uptime and performance
Malvern Panalytical instruments are built to last and good maintenance and care ensure continued peak performance for many years to come. Service Agreements build this into your schedule and budget, based on preventive maintenance and remote support . Different levels include repair visits, response times and parts coverage as needed. OQ services, training and application consultancy can be added as well.
|
|
|
|
Advanced analysis and actionable insights
|
|
|
|
Easy start and streamlined routine operations
|
|
|
|
Want to speak to a service expert or find the services that are just right for you?
Reach out to our experts and evaluate your services needs and business risks by answering a few questions around the usage of your Malvern Panalytical instrument.
|
|
|
*Please note that the exact details of our services and their availability are determined by a variety of customer-specific factors, such as product type, product configuration, or location. |
||

Powerful characterization tools for characterization of the thermal stability of proteins and other biomolecules in biopharmaceutical development, manufacture and beyond. Simple to use, requires little assay development, and needs no labeling or immobilization.