Label-free interaction analysis
Label-free interaction analysis techniques for the direct study of native molecules such as proteins, without using artificial probes or labels
The Kinetics Guide | Binding kinetics with the WAVE system. Download now
Download nowLabel-free interaction analysis techniques for the direct study of native molecules such as proteins, without using artificial probes or labels

Traditional label-based approaches often require individual bioassay components to be labeled for detection to be possible. Many conventional bioassays use an immobilized ligand to capture a target analyte from solution, before employing specialized detection reagents to analyze the interaction between the two molecules.
Examples of these approaches include as enzyme-linked immunosorbent assay (ELISA) or bead-based flow cytometry. Labeling can alter the structure and/or function of biomolecules and skew experimental results.
In some cases, they can completely deactivate the biomolecule that is being studied. Moreover, the non-specific binding of the label components to other bioassay components can produce unwanted background signal. This is especially problematic when working with complex samples like cell culture supernatants, serum, or plasma.
Label-free quantification of molecular interactions presents an attractive alternative to traditional label-based approaches. Label-free technology (also label-free detection or label-free sensing) uses impedance or optical biosensors to measure the changes that occur upon analyte binding to a ligand immobilized on a biosensor surface. This approach enables the interaction to be monitored in real-time without artificial manipulation of the individual assay components.
Get an overview of how label-free technology and innovative GCI technology differs from traditional bioassay techniques below.
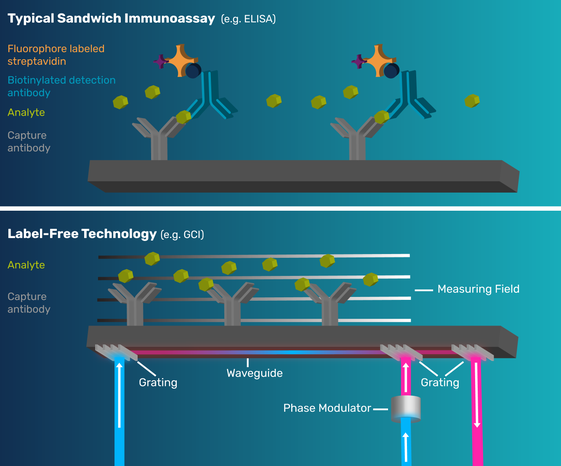
Label-free interaction analysis techniques are used in the direct study of native molecules. They generate biologically relevant data that enable the understanding of molecular interactions without using artificial probes or labels and therefore provide data that is more likely to reflect the natural behavior of these molecules. This is key to providing a robust body of knowledge to help develop a good understanding of structure activity relationships, central disease research and to discover safe and efficacious drugs.
Importantly, label-free technology allows researchers to carry out analyses that are simply not possible using conventional methods that provide only end-point results. These include kinetic and affinity analysis, and evaluation of binding specificity, which are enabled by the dynamic, real-time interaction analysis afforded by label-free monitoring.
Since common additives to protein solutions can interfere with the detection of fluorescent labels, label-free assays typically work in most biological buffers. Label-free methods are also desirable because they are typically compatible with real-time detection. Label-free technologies are attractive to researchers across a diverse range of applications, such as: biological research, biotherapeutic and small molecule drug discovery and development, immunogenicity studies, and vaccine development.
Several label-free approaches for the detection of biomolecular interactions are currently available. These include the use of optical biosensors, such as GCI, diffraction, quartz crystal microbalance and microcalorimetry.
Malvern Panalytical offers the WAVEsystem biosensor and the MicroCal PEAQ-ITC and Microcal PEAQ-DSC microcalorimeters - powerful and well-established label-free, in-solution analysis instruments.

Next-gen bioanalysis platform
The WAVEsystem biosensors use GCI technology to establish critical reaction parameters such as the association rate constant (ka), dissociation rate constant (kd), and the equilibrium dissociation constant (KD) of biomolecular interactions in real-time.
The resulting information-rich data provides insight into the biological processes and the mechanisms driving intra- and intermolecular interactions. This label-free technology allows native ligand to be immobilized on a biosensor surface.

MicroCal PEAQ-ITC delivers exceptional sensitivity and high quality data wi...
MicroCal PEAQ- ITC can detect the association and dissociation of molecular complexes by measuring changes in the heat released or absorbed when molecules interact.
The resulting information-rich data provides insight into the biological processes and the mechanisms driving intra- and intermolecular interactions.

Differential scanning calorimeters
Microcal PEAQ-DSC can detect the heat changes that occur when a macromolecule, such as a protein or nucleic acid is unfolded or a liposomal micelle dissociates.
This is often used to find stabilizing conditions for biologics, for assay development for drug screening and protein characterization and for vaccine development.